Helicase-like functions in phosphate loop containing beta-alpha polypeptides
- PMID: 33846247
- PMCID: PMC8072362
- DOI: 10.1073/pnas.2016131118
Helicase-like functions in phosphate loop containing beta-alpha polypeptides
Abstract
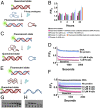
The P-loop Walker A motif underlies hundreds of essential enzyme families that bind nucleotide triphosphates (NTPs) and mediate phosphoryl transfer (P-loop NTPases), including the earliest DNA/RNA helicases, translocases, and recombinases. What were the primordial precursors of these enzymes? Could these large and complex proteins emerge from simple polypeptides? Previously, we showed that P-loops embedded in simple βα repeat proteins bind NTPs but also, unexpectedly so, ssDNA and RNA. Here, we extend beyond the purely biophysical function of ligand binding to demonstrate rudimentary helicase-like activities. We further constructed simple 40-residue polypeptides comprising just one β-(P-loop)-α element. Despite their simplicity, these P-loop prototypes confer functions such as strand separation and exchange. Foremost, these polypeptides unwind dsDNA, and upon addition of NTPs, or inorganic polyphosphates, release the bound ssDNA strands to allow reformation of dsDNA. Binding kinetics and low-resolution structural analyses indicate that activity is mediated by oligomeric forms spanning from dimers to high-order assemblies. The latter are reminiscent of extant P-loop recombinases such as RecA. Overall, these P-loop prototypes compose a plausible description of the sequence, structure, and function of the earliest P-loop NTPases. They also indicate that multifunctionality and dynamic assembly were key in endowing short polypeptides with elaborate, evolutionarily relevant functions.
Keywords: P-loop; Walker A; multifunctionality; polyphosphate; protein evolution.
Conflict of interest statement
The authors declare no competing interest.
Figures




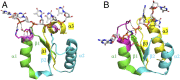
Similar articles
-
Simple yet functional phosphate-loop proteins.Proc Natl Acad Sci U S A. 2018 Dec 18;115(51):E11943-E11950. doi: 10.1073/pnas.1812400115. Epub 2018 Nov 30. Proc Natl Acad Sci U S A. 2018. PMID: 30504143 Free PMC article.
-
A Distinct Motif in a Prokaryotic Small Ras-Like GTPase Highlights Unifying Features of Walker B Motifs in P-Loop NTPases.J Mol Biol. 2020 Sep 18;432(20):5544-5564. doi: 10.1016/j.jmb.2020.07.024. Epub 2020 Aug 1. J Mol Biol. 2020. PMID: 32750390
-
STAND, a class of P-loop NTPases including animal and plant regulators of programmed cell death: multiple, complex domain architectures, unusual phyletic patterns, and evolution by horizontal gene transfer.J Mol Biol. 2004 Oct 8;343(1):1-28. doi: 10.1016/j.jmb.2004.08.023. J Mol Biol. 2004. PMID: 15381417 Review.
-
On the Origins of Enzymes: Phosphate-Binding Polypeptides Mediate Phosphoryl Transfer to Synthesize Adenosine Triphosphate.J Am Chem Soc. 2023 Mar 23;145(15):8344-54. doi: 10.1021/jacs.2c08636. Online ahead of print. J Am Chem Soc. 2023. PMID: 36951643 Free PMC article.
-
Unraveling DNA helicases. Motif, structure, mechanism and function.Eur J Biochem. 2004 May;271(10):1849-63. doi: 10.1111/j.1432-1033.2004.04094.x. Eur J Biochem. 2004. PMID: 15128295 Review.
Cited by
-
Enzyme catalysis prior to aromatic residues: Reverse engineering of a dephospho-CoA kinase.Protein Sci. 2021 May;30(5):1022-1034. doi: 10.1002/pro.4068. Epub 2021 Mar 26. Protein Sci. 2021. PMID: 33739538 Free PMC article.
-
Why we are made of proteins and nucleic acids: Structural biology views on extraterrestrial life.Biophys Physicobiol. 2023 Jun 2;20(2):e200026. doi: 10.2142/biophysico.bppb-v20.0026. eCollection 2023. Biophys Physicobiol. 2023. PMID: 38496239 Free PMC article.
-
Back in time to the Gly-rich prototype of the phosphate binding elementary function.Curr Res Struct Biol. 2024 Apr 9;7:100142. doi: 10.1016/j.crstbi.2024.100142. eCollection 2024. Curr Res Struct Biol. 2024. PMID: 38655428 Free PMC article. Review.
-
Distribution of Polyphosphate Kinase 2 Genes in Bacteria Underscores a Dynamic Evolutionary History.Proteins. 2025 May;93(5):972-980. doi: 10.1002/prot.26780. Epub 2024 Dec 18. Proteins. 2025. PMID: 39691974 Free PMC article.
-
On Protein Loops, Prior Molecular States and Common Ancestors of Life.J Mol Evol. 2024 Oct;92(5):624-646. doi: 10.1007/s00239-024-10167-y. Epub 2024 Apr 23. J Mol Evol. 2024. PMID: 38652291 Free PMC article. Review.
References
-
- Bleichert F., Botchan M. R., Berger J. M., Mechanisms for initiating cellular DNA replication. Science 355, eaah6317 (2017). - PubMed
-
- Eck R. V., Dayhoff M. O., Evolution of the structure of ferredoxin based on living relics of primitive amino acid sequences. Science 152, 363–366 (1966). - PubMed
-
- Romero M. L. R., Rabin A., Tawfik D. S., Functional proteins from short peptides : Dayhoff s hypothesis turns 50. Angew. Chem. Int. Ed. Engl. 1980, 15966–15971 (2016). - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

