The Botrytis cinerea Crh1 transglycosylase is a cytoplasmic effector triggering plant cell death and defense response
- PMID: 33846308
- PMCID: PMC8042016
- DOI: 10.1038/s41467-021-22436-1
The Botrytis cinerea Crh1 transglycosylase is a cytoplasmic effector triggering plant cell death and defense response
Abstract
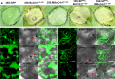
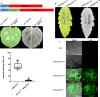
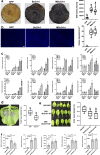
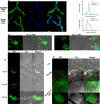
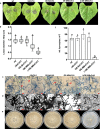
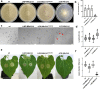
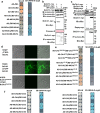
Crh proteins catalyze crosslinking of chitin and glucan polymers in fungal cell walls. Here, we show that the BcCrh1 protein from the phytopathogenic fungus Botrytis cinerea acts as a cytoplasmic effector and elicitor of plant defense. BcCrh1 is localized in vacuoles and the endoplasmic reticulum during saprophytic growth. However, upon plant infection, the protein accumulates in infection cushions; it is then secreted to the apoplast and translocated into plant cells, where it induces cell death and defense responses. Two regions of 53 and 35 amino acids are sufficient for protein uptake and cell death induction, respectively. BcCrh1 mutant variants that are unable to dimerize lack transglycosylation activity, but are still able to induce plant cell death. Furthermore, Arabidopsis lines expressing the bccrh1 gene exhibit reduced sensitivity to B. cinerea, suggesting a potential use of the BcCrh1 protein in plant immunization against this necrotrophic pathogen.
Conflict of interest statement
The authors declare no competing interests.
Figures

Similar articles
-
A LysM Effector Mediates Adhesion and Plant Immunity Suppression in the Necrotrophic Fungus Botrytis cinerea.J Basic Microbiol. 2025 May;65(5):e2400552. doi: 10.1002/jobm.202400552. Epub 2024 Dec 10. J Basic Microbiol. 2025. PMID: 39655398
-
The novel elicitor AsES triggers a defense response against Botrytis cinerea in Arabidopsis thaliana.Plant Sci. 2015 Dec;241:120-7. doi: 10.1016/j.plantsci.2015.09.025. Epub 2015 Oct 9. Plant Sci. 2015. PMID: 26706064
-
BcIEB1, a Botrytis cinerea secreted protein, elicits a defense response in plants.Plant Sci. 2016 Sep;250:115-124. doi: 10.1016/j.plantsci.2016.06.009. Epub 2016 Jun 14. Plant Sci. 2016. PMID: 27457989
-
Botrytis small RNA Bc-siR37 suppresses plant defense genes by cross-kingdom RNAi.RNA Biol. 2017 Apr 3;14(4):421-428. doi: 10.1080/15476286.2017.1291112. Epub 2017 Mar 7. RNA Biol. 2017. PMID: 28267415 Free PMC article. Review.
-
Recent Advances in the Study of the Plant Pathogenic Fungus Botrytis cinerea and its Interaction with the Environment.Curr Protein Pept Sci. 2017;18(10):976-989. doi: 10.2174/1389203717666160809160915. Curr Protein Pept Sci. 2017. PMID: 27526927 Review.
Cited by
-
Genome-wide identification of long non-coding RNA for Botrytis cinerea during infection to tomato (Solanum lycopersicum) leaves.BMC Genomics. 2025 Jan 6;26(1):7. doi: 10.1186/s12864-024-11171-8. BMC Genomics. 2025. PMID: 39762752 Free PMC article.
-
Effectors of Plant Necrotrophic Fungi.Front Plant Sci. 2021 Jun 4;12:687713. doi: 10.3389/fpls.2021.687713. eCollection 2021. Front Plant Sci. 2021. PMID: 34149788 Free PMC article. Review.
-
Comprehensive Genomic and Proteomic Analysis Identifies Effectors of Fusarium oxysporum f. sp. melongenae.J Fungi (Basel). 2024 Nov 28;10(12):828. doi: 10.3390/jof10120828. J Fungi (Basel). 2024. PMID: 39728324 Free PMC article.
-
Secreted Glycoside Hydrolase Proteins as Effectors and Invasion Patterns of Plant-Associated Fungi and Oomycetes.Front Plant Sci. 2022 Mar 10;13:853106. doi: 10.3389/fpls.2022.853106. eCollection 2022. Front Plant Sci. 2022. PMID: 35360318 Free PMC article. Review.
-
Effectors of plants pathogenic fungi and fungal like microbes: a comprehensive review on mechanisms, roles, and host interactions.Front Plant Sci. 2025 Jul 29;16:1626960. doi: 10.3389/fpls.2025.1626960. eCollection 2025. Front Plant Sci. 2025. PMID: 40799271 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Other Literature Sources

