Concerted control of the LaRAV1-LaCDKB1;3 module by temperature during dormancy release and reactivation of larch
- PMID: 33847364
- PMCID: PMC8498939
- DOI: 10.1093/treephys/tpab052
Concerted control of the LaRAV1-LaCDKB1;3 module by temperature during dormancy release and reactivation of larch
Abstract
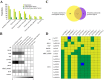
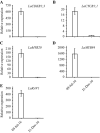
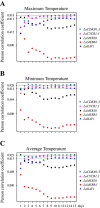
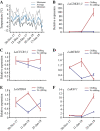
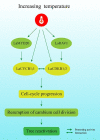
Dormancy release and reactivation of temperate-zone trees involve the temperature-modulated expression of cell-cycle genes. However, information on the detailed regulatory mechanism is limited. Here, we compared the transcriptomes of the stems of active and dormant larch trees, emphasizing the expression patterns of cell-cycle genes and transcription factors and assessed their relationships and responses to temperatures. Twelve cell-cycle genes and 31 transcription factors were strongly expressed in the active stage. Promoter analysis suggested that these 12 genes might be regulated by transcription factors from 10 families. Altogether, 73 cases of regulation between 16 transcription factors and 12 cell-cycle genes were predicted, while the regulatory interactions between LaMYB20 and LaCYCB1;1, and LaRAV1 and LaCDKB1;3 were confirmed by yeast one-hybrid and dual-luciferase assays. Last, we found that LaRAV1 and LaCDKB1;3 had almost the same expression patterns during dormancy release and reactivation induced naturally or artificially by temperature, indicating that the LaRAV1-LaCDKB1;3 module functions in the temperature-modulated dormancy release and reactivation of larch trees. These results provide new insights into the link between temperature and cell-cycle gene expression, helping to understand the temperature control of tree growth and development in the context of climate change.
Keywords: cambium; cell-cycle gene; dormancy release; larch; reactivation; temperature; transcription factor.
© The Author(s) 2021. Published by Oxford University Press. All rights reserved. For permissions, please e-mail: journals.permission@oup.com.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures






Similar articles
-
Cambium Reactivation Is Closely Related to the Cell-Cycle Gene Configuration in Larix kaempferi.Int J Mol Sci. 2024 Mar 22;25(7):3578. doi: 10.3390/ijms25073578. Int J Mol Sci. 2024. PMID: 38612390 Free PMC article.
-
Over-Expression of the Cell-Cycle Gene LaCDKB1;2 Promotes Cell Proliferation and the Formation of Normal Cotyledonary Embryos during Larix kaempferi Somatic Embryogenesis.Genes (Basel). 2021 Sep 17;12(9):1435. doi: 10.3390/genes12091435. Genes (Basel). 2021. PMID: 34573419 Free PMC article.
-
Transcriptome profiles revealed molecular mechanisms of alternating temperatures in breaking the epicotyl morphophysiological dormancy of Polygonatum sibiricum seeds.BMC Plant Biol. 2021 Aug 12;21(1):370. doi: 10.1186/s12870-021-03147-7. BMC Plant Biol. 2021. PMID: 34384392 Free PMC article.
-
Photoperiod- and temperature-mediated control of phenology in trees - a molecular perspective.New Phytol. 2017 Jan;213(2):511-524. doi: 10.1111/nph.14346. Epub 2016 Nov 30. New Phytol. 2017. PMID: 27901272 Review.
-
I Want to (Bud) Break Free: The Potential Role of DAM and SVP-Like Genes in Regulating Dormancy Cycle in Temperate Fruit Trees.Front Plant Sci. 2019 Jan 10;9:1990. doi: 10.3389/fpls.2018.01990. eCollection 2018. Front Plant Sci. 2019. PMID: 30687377 Free PMC article. Review.
Cited by
-
Screening and Functional Evaluation of Four Larix kaempferi Promoters.Plants (Basel). 2024 Oct 3;13(19):2777. doi: 10.3390/plants13192777. Plants (Basel). 2024. PMID: 39409647 Free PMC article.
-
Exploring the Seasonal Dynamics and Molecular Mechanism of Wood Formation in Gymnosperm Trees.Int J Mol Sci. 2023 May 11;24(10):8624. doi: 10.3390/ijms24108624. Int J Mol Sci. 2023. PMID: 37239969 Free PMC article. Review.
-
Cambium Reactivation Is Closely Related to the Cell-Cycle Gene Configuration in Larix kaempferi.Int J Mol Sci. 2024 Mar 22;25(7):3578. doi: 10.3390/ijms25073578. Int J Mol Sci. 2024. PMID: 38612390 Free PMC article.
-
Over-Expression of the Cell-Cycle Gene LaCDKB1;2 Promotes Cell Proliferation and the Formation of Normal Cotyledonary Embryos during Larix kaempferi Somatic Embryogenesis.Genes (Basel). 2021 Sep 17;12(9):1435. doi: 10.3390/genes12091435. Genes (Basel). 2021. PMID: 34573419 Free PMC article.
-
VS-Cambium-Developer: A New Predictive Model of Cambium Functioning under the Influence of Environmental Factors.Plants (Basel). 2023 Oct 17;12(20):3594. doi: 10.3390/plants12203594. Plants (Basel). 2023. PMID: 37896057 Free PMC article.
References
-
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410. - PubMed
-
- Anh Tuan P, Bai S, Saito T, Imai T, Ito A, Moriguchi T (2016) Involvement of EARLY BUD-BREAK, an AP2/ERF transcription factor gene, in bud break in Japanese pear (Pyrus pyrifolia Nakai) lateral flower buds: expression, histone modifications and possible target genes. Plant Cell Physiol 57:1038–1047. - PubMed
-
- Begum S, Kudo K, Rahman MH et al. (2018) Climate change and the regulation of wood formation in trees by temperature. Trees 32:3–15.
-
- Berckmans B, De Veylder L (2009) Transcriptional control of the cell cycle. Curr Opin Plant Biol 12:599–605. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

