Dynamics of the Bacillus subtilis Min System
- PMID: 33849976
- PMCID: PMC8092234
- DOI: 10.1128/mBio.00296-21
Dynamics of the Bacillus subtilis Min System
Abstract
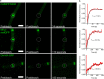
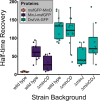
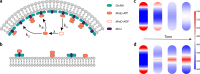
Division site selection is a vital process to ensure generation of viable offspring. In many rod-shaped bacteria, a dynamic protein system, termed the Min system, acts as a central regulator of division site placement. The Min system is best studied in Escherichia coli, where it shows a remarkable oscillation from pole to pole with a time-averaged density minimum at midcell. Several components of the Min system are conserved in the Gram-positive model organism Bacillus subtilis However, in B. subtilis, it is commonly believed that the system forms a stationary bipolar gradient from the cell poles to midcell. Here, we show that the Min system of B. subtilis localizes dynamically to active sites of division, often organized in clusters. We provide physical modeling using measured diffusion constants that describe the observed enrichment of the Min system at the septum. Mathematical modeling suggests that the observed localization pattern of Min proteins corresponds to a dynamic equilibrium state. Our data provide evidence for the importance of ongoing septation for the Min dynamics, consistent with a major role of the Min system in controlling active division sites but not cell pole areas.IMPORTANCE The molecular mechanisms that help to place the division septum in bacteria is of fundamental importance to ensure cell proliferation and maintenance of cell shape and size. The Min protein system, found in many rod-shaped bacteria, is thought to play a major role in division site selection. It was assumed that there are strong differences in the functioning and in the dynamics of the Min system in E. coli and B. subtilis Most previous attempts to address Min protein dynamics in B. subtilis have been hampered by the use of overexpression constructs. Here, functional fusions to Min proteins have been constructed by allelic exchange and state-of-the-art imaging techniques allowed to unravel an unexpected fast dynamic behavior of the B. subtilis Min system. Our data show that the molecular mechanisms leading to Min protein dynamics are not fundamentally different in E. coli and B. subtilis.
Keywords: B. subtilis; FRAP; Min system; PALM; cell division; protein patterns; reaction diffusion equations; super resolution microscopy.
Copyright © 2021 Feddersen et al.
Figures


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous
