Prediction of single-cell mechanisms for disease progression in hypertrophic remodelling by a trans-omics approach
- PMID: 33854108
- PMCID: PMC8047020
- DOI: 10.1038/s41598-021-86821-y
Prediction of single-cell mechanisms for disease progression in hypertrophic remodelling by a trans-omics approach
Abstract
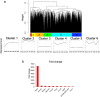
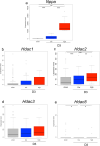
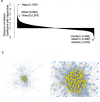
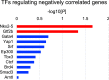
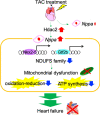
Heart failure is a heterogeneous disease with multiple risk factors and various pathophysiological types, which makes it difficult to understand the molecular mechanisms involved. In this study, we proposed a trans-omics approach for predicting molecular pathological mechanisms of heart failure and identifying marker genes to distinguish heterogeneous phenotypes, by integrating multiple omics data including single-cell RNA-seq, ChIP-seq, and gene interactome data. We detected a significant increase in the expression level of natriuretic peptide A (Nppa), after stress loading with transverse aortic constriction (TAC), and showed that cardiomyocytes with high Nppa expression displayed specific gene expression patterns. Multiple NADH ubiquinone complex family, which are associated with the mitochondrial electron transport system, were negatively correlated with Nppa expression during the early stages of cardiac hypertrophy. Large-scale ChIP-seq data analysis showed that Nkx2-5 and Gtf2b were transcription factors characteristic of high-Nppa-expressing cardiomyocytes. Nppa expression levels may, therefore, represent a useful diagnostic marker for heart failure.
Conflict of interest statement
The authors declare no competing interests.
Figures




References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

