Glioblastomas acquire myeloid-affiliated transcriptional programs via epigenetic immunoediting to elicit immune evasion
- PMID: 33857425
- PMCID: PMC8099351
- DOI: 10.1016/j.cell.2021.03.023
Glioblastomas acquire myeloid-affiliated transcriptional programs via epigenetic immunoediting to elicit immune evasion
Abstract
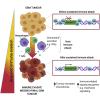
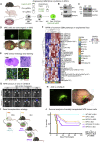
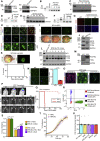
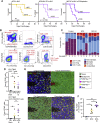
Glioblastoma multiforme (GBM) is an aggressive brain tumor for which current immunotherapy approaches have been unsuccessful. Here, we explore the mechanisms underlying immune evasion in GBM. By serially transplanting GBM stem cells (GSCs) into immunocompetent hosts, we uncover an acquired capability of GSCs to escape immune clearance by establishing an enhanced immunosuppressive tumor microenvironment. Mechanistically, this is not elicited via genetic selection of tumor subclones, but through an epigenetic immunoediting process wherein stable transcriptional and epigenetic changes in GSCs are enforced following immune attack. These changes launch a myeloid-affiliated transcriptional program, which leads to increased recruitment of tumor-associated macrophages. Furthermore, we identify similar epigenetic and transcriptional signatures in human mesenchymal subtype GSCs. We conclude that epigenetic immunoediting may drive an acquired immune evasion program in the most aggressive mesenchymal GBM subtype by reshaping the tumor immune microenvironment.
Keywords: DNA methylation; chemokine; epigenetics; glioblastoma; immune evasion; immunoediting; interferon signaling; macrophage; neural stem cell; syngeneic.
Copyright © 2021 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests S.M.P is a co-founder and shareholder of Cellinta., a biotechnology start-up that is developing cancer therapeutics, including for glioblastoma, and acts as an advisor to the company. J.W.P is a co-founder and shareholder of Macomics. S.A.Q. is co-founder and shareholder and Chief Scientific Officer for Achilles Therapeutics. The other authors declare no competing interests.
Figures
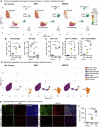





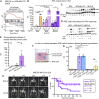


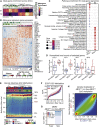

Comment in
-
The imitation game: How glioblastoma outmaneuvers immune attack.Cell. 2021 Apr 29;184(9):2278-2281. doi: 10.1016/j.cell.2021.04.008. Cell. 2021. PMID: 33930294
References
-
- Benjamini Y., Hochberg Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. Ser. A Stat. Soc. 1995;57:289–300.
Publication types
MeSH terms
Substances
Grants and funding
- 17368/CRUK_/Cancer Research UK/United Kingdom
- 20837/CRUK_/Cancer Research UK/United Kingdom
- MC_UU_00009/2/MRC_/Medical Research Council/United Kingdom
- A21922/CRUK_/Cancer Research UK/United Kingdom
- 27590/CRUK_/Cancer Research UK/United Kingdom
- 22246/CRUK_/Cancer Research UK/United Kingdom
- 20764/CRUK_/Cancer Research UK/United Kingdom
- 21203/CRUK_/Cancer Research UK/United Kingdom
- 23615/CRUK_/Cancer Research UK/United Kingdom
- 23459/CRUK_/Cancer Research UK/United Kingdom
- A28592/CRUK_/Cancer Research UK/United Kingdom
- 21992/CRUK_/Cancer Research UK/United Kingdom
- 28592/CRUK_/Cancer Research UK/United Kingdom
- A17368/CRUK_/Cancer Research UK/United Kingdom
- C39669/A25919/CRUK_/Cancer Research UK/United Kingdom
- 25919/CRUK_/Cancer Research UK/United Kingdom
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

