Novel, non-invasive markers for detecting therapy induced neuroendocrine differentiation in castration-resistant prostate cancer patients
- PMID: 33859239
- PMCID: PMC8050049
- DOI: 10.1038/s41598-021-87441-2
Novel, non-invasive markers for detecting therapy induced neuroendocrine differentiation in castration-resistant prostate cancer patients
Abstract
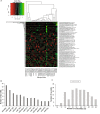
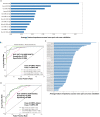
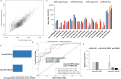
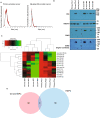
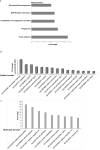
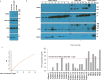
Neuroendocrine prostate cancer (NEPC), a highly aggressive variant of castration-resistant prostate cancer (CRPC), often emerges upon treatment with androgen pathway inhibitors, via neuroendocrine differentiation. Currently, NEPC diagnosis is challenging as available markers are not sufficiently specific. Our objective was to identify novel, extracellular vesicles (EV)-based biomarkers for diagnosing NEPC. Towards this, we performed small RNA next generation sequencing in serum EVs isolated from a cohort of CRPC patients with adenocarcinoma characteristics (CRPC-Adeno) vs CRPC-NE and identified significant dysregulation of 182 known and 4 novel miRNAs. We employed machine learning algorithms to develop an 'EV-miRNA classifier' that could robustly stratify 'CRPC-NE' from 'CRPC-Adeno'. Examination of protein repertoire of exosomes from NEPC cellular models by mass spectrometry identified thrombospondin 1 (TSP1) as a specific biomarker. In view of our results, we propose that a miRNA panel and TSP1 can be used as novel, non-invasive tools to identify NEPC and guide treatment decisions. In conclusion, our study identifies for the first time, novel non-invasive exosomal/extracellular vesicle based biomarkers for detecting neuroendocrine differentiation in advanced castration resistant prostate cancer patients with important translational implications in clinical management of these patients that is currently extremely challenging.
Conflict of interest statement
The authors declare no competing interests.
Figures

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

