Moonlighting of mitotic regulators in cilium disassembly
- PMID: 33860332
- PMCID: PMC8233288
- DOI: 10.1007/s00018-021-03827-5
Moonlighting of mitotic regulators in cilium disassembly
Abstract
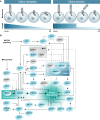
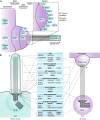
Correct timing of cellular processes is essential during embryological development and to maintain the balance between healthy proliferation and tumour formation. Assembly and disassembly of the primary cilium, the cell's sensory signalling organelle, are linked to cell cycle timing in the same manner as spindle pole assembly and chromosome segregation. Mitotic processes, ciliary assembly, and ciliary disassembly depend on the centrioles as microtubule-organizing centres (MTOC) to regulate polymerizing and depolymerizing microtubules. Subsequently, other functional protein modules are gathered to potentiate specific protein-protein interactions. In this review, we show that a significant subset of key mitotic regulator proteins is moonlighting at the cilium, among which PLK1, AURKA, CDC20, and their regulators. Although ciliary assembly defects are linked to a variety of ciliopathies, ciliary disassembly defects are more often linked to brain development and tumour formation. Acquiring a better understanding of the overlap in regulators of ciliary disassembly and mitosis is essential in finding therapeutic targets for the different diseases and types of tumours associated with these regulators.
Keywords: Cell cycle regulators; Centrioles; Cilia; Cilium resorption; Tumour development; WNT.
Conflict of interest statement
The authors declare they have no conflict of interest.
Figures

Similar articles
-
PCM1 recruits Plk1 to the pericentriolar matrix to promote primary cilia disassembly before mitotic entry.J Cell Sci. 2013 Mar 15;126(Pt 6):1355-65. doi: 10.1242/jcs.114918. Epub 2013 Jan 23. J Cell Sci. 2013. PMID: 23345402
-
The regulation of cilium assembly and disassembly in development and disease.Development. 2018 Sep 17;145(18):dev151407. doi: 10.1242/dev.151407. Development. 2018. PMID: 30224385 Free PMC article. Review.
-
Primary cilia biogenesis and associated retinal ciliopathies.Semin Cell Dev Biol. 2021 Feb;110:70-88. doi: 10.1016/j.semcdb.2020.07.013. Epub 2020 Jul 31. Semin Cell Dev Biol. 2021. PMID: 32747192 Free PMC article. Review.
-
Distinct roles of centriole distal appendage proteins in ciliary assembly and disassembly.Cell Commun Signal. 2024 Dec 18;22(1):607. doi: 10.1186/s12964-024-01962-7. Cell Commun Signal. 2024. PMID: 39696441 Free PMC article.
-
The primary cilium as a multiple cellular signaling scaffold in development and disease.BMB Rep. 2012 Aug;45(8):427-32. doi: 10.5483/BMBRep.2012.45.8.167. BMB Rep. 2012. PMID: 22917026
Cited by
-
CDC20-Mediated Selective Autophagy Degradation of PBRM1 Affects Immunotherapy for Renal Cell Carcinoma.Adv Sci (Weinh). 2025 Feb;12(5):e2412967. doi: 10.1002/advs.202412967. Epub 2024 Dec 10. Adv Sci (Weinh). 2025. PMID: 39656940 Free PMC article.
-
Insights into the Regulation of Ciliary Disassembly.Cells. 2021 Nov 1;10(11):2977. doi: 10.3390/cells10112977. Cells. 2021. PMID: 34831200 Free PMC article. Review.
-
CDC20 in and out of mitosis: a prognostic factor and therapeutic target in hematological malignancies.J Exp Clin Cancer Res. 2022 Apr 30;41(1):159. doi: 10.1186/s13046-022-02363-9. J Exp Clin Cancer Res. 2022. PMID: 35490245 Free PMC article. Review.
-
Primary Cilium Is Involved in Stem Cell Differentiation and Renewal through the Regulation of Multiple Signaling Pathways.Cells. 2021 Jun 8;10(6):1428. doi: 10.3390/cells10061428. Cells. 2021. PMID: 34201019 Free PMC article. Review.
-
Chlamydia trachomatis induces disassembly of the primary cilium to promote the intracellular infection.PLoS Pathog. 2024 Jun 17;20(6):e1012303. doi: 10.1371/journal.ppat.1012303. eCollection 2024 Jun. PLoS Pathog. 2024. PMID: 38885287 Free PMC article.
References
-
- Ross AJ, May-Simera H, Eichers ER, Kai M, Hill J, Jagger DJ, Leitch CC, Chapple JP, Munro PM, Fisher S, Tan PL, Phillips HM, Leroux MR, Henderson DJ, Murdoch JN, Copp AJ, Eliot MM, Lupski JR, Kemp DT, Dollfus H, Tada M, Katsanis N, Forge A, Beales PL. Disruption of Bardet-Biedl syndrome ciliary proteins perturbs planar cell polarity in vertebrates. Nat Genet. 2005;37(10):1135–1140. doi: 10.1038/ng1644. - DOI - PubMed
-
- Simons M, Gloy J, Ganner A, Bullerkotte A, Bashkurov M, Kronig C, Schermer B, Benzing T, Cabello OA, Jenny A, Mlodzik M, Polok B, Driever W, Obara T, Walz G. Inversin, the gene product mutated in nephronophthisis type II, functions as a molecular switch between Wnt signaling pathways. Nat Genet. 2005;37(5):537–543. doi: 10.1038/ng1552. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

