Design of multi-scale protein complexes by hierarchical building block fusion
- PMID: 33863889
- PMCID: PMC8052403
- DOI: 10.1038/s41467-021-22276-z
Design of multi-scale protein complexes by hierarchical building block fusion
Abstract
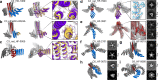
A systematic and robust approach to generating complex protein nanomaterials would have broad utility. We develop a hierarchical approach to designing multi-component protein assemblies from two classes of modular building blocks: designed helical repeat proteins (DHRs) and helical bundle oligomers (HBs). We first rigidly fuse DHRs to HBs to generate a large library of oligomeric building blocks. We then generate assemblies with cyclic, dihedral, and point group symmetries from these building blocks using architecture guided rigid helical fusion with new software named WORMS. X-ray crystallography and cryo-electron microscopy characterization show that the hierarchical design approach can accurately generate a wide range of assemblies, including a 43 nm diameter icosahedral nanocage. The computational methods and building block sets described here provide a very general route to de novo designed protein nanomaterials.
Conflict of interest statement
The authors declare no competing interests.
Figures




References
Publication types
MeSH terms
Substances
Grants and funding
- R01 GM120553/GM/NIGMS NIH HHS/United States
- P30 GM124169/GM/NIGMS NIH HHS/United States
- S10 OD023476/OD/NIH HHS/United States
- HHMI/Howard Hughes Medical Institute/United States
- S10 OD018483/OD/NIH HHS/United States
- S10 RR029300/RR/NCRR NIH HHS/United States
- P30 GM124165/GM/NIGMS NIH HHS/United States
- S10 OD019994/OD/NIH HHS/United States
- T32 GM008268/GM/NIGMS NIH HHS/United States
- HHSN272201700059C/AI/NIAID NIH HHS/United States
- P41 GM103310/GM/NIGMS NIH HHS/United States
- S10 OD021527/OD/NIH HHS/United States
- T32 GM007270/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

