HIV-1 Capsid Core: A Bullet to the Heart of the Target Cell
- PMID: 33868211
- PMCID: PMC8046902
- DOI: 10.3389/fmicb.2021.652486
HIV-1 Capsid Core: A Bullet to the Heart of the Target Cell
Abstract
The first step of the intracellular phase of retroviral infection is the release of the viral capsid core in the cytoplasm. This structure contains the viral genetic material that will be reverse transcribed and integrated into the genome of infected cells. Up to recent times, the role of the capsid core was considered essentially to protect this genetic material during the earlier phases of this process. However, increasing evidence demonstrates that the permanence inside the cell of the capsid as an intact, or almost intact, structure is longer than thought. This suggests its involvement in more aspects of the infectious cycle than previously foreseen, particularly in the steps of viral genomic material translocation into the nucleus and in the phases preceding integration. During the trip across the infected cell, many host factors are brought to interact with the capsid, some possessing antiviral properties, others, serving as viral cofactors. All these interactions rely on the properties of the unique component of the capsid core, the capsid protein CA. Likely, the drawback of ensuring these multiple functions is the extreme genetic fragility that has been shown to characterize this protein. Here, we recapitulate the busy agenda of an HIV-1 capsid in the infectious process, in particular in the light of the most recent findings.
Keywords: HIV-1; capsid; cellular cofactors; genetic fragility; nuclear transport; restriction factors; reverse transcription; uncoating.
Copyright © 2021 Toccafondi, Lener and Negroni.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


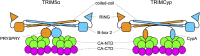
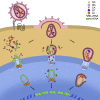
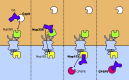
References
-
- Accola M. A., Strack B., Göttlinger H. G. (2000). Efficient particle production by minimal Gag constructs which retain the carboxy-terminal domain of human immunodeficiency virus type 1 Capsid-p2 and a late assembly domain. J. Virol. 74 5395–5402. 10.1128/jvi.74.12.5395-5402.2000 - DOI - PMC - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources

