Alteration of the Gut Microbiome in Chronic Kidney Disease Patients and Its Association With Serum Free Immunoglobulin Light Chains
- PMID: 33868230
- PMCID: PMC8047322
- DOI: 10.3389/fimmu.2021.609700
Alteration of the Gut Microbiome in Chronic Kidney Disease Patients and Its Association With Serum Free Immunoglobulin Light Chains
Abstract
Objectives: Gut dysbiosis is associated with chronic kidney disease (CKD), and serum free immunoglobulin light chains (FLCs) are biomarkers for CKD. This study aims to assess the CKD gut microbiome and to determine its impact on serum FLC levels.
Methods: To control for confounders, 100 patients and sex- and age-matched healthy controls (HCs) were recruited. The gut microbiome was assessed by sequencing 16S rRNA gene V3-V4 hypervariable regions. Phylogenetic Investigation of Communities by Reconstruction of Unobserved States was applied to infer functional metabolic pathways. When observing group differences in the microbiome and predicted metabolic pathways, demographic confounders were adjusted using binary logistic regression; when examining impacts of the gut microbiome and metabolic pathways on serum FLCs, factors influencing FLC levels were adjusted using multiple regression.
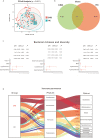
Results: Principal coordinate analysis revealed a significantly different bacterial community between the CKD and HC groups (P < 0.05). After adjusting for confounders, lower Chao 1, observed species and Shannon indices based on binary logistic regression predicted CKD prevalence. Actinobacteria, Alistipes, Bifidobacterium and Bifidobacterium longum enrichment, upregulation of metabolic pathways of bacterial toxin, chloroalkane and chloroalkene degradation, and Staphylococcus aureus infection also predicted CKD prevalence (P < 0.05). Furthermore, depletion of Actinobacteria and Bifidobacterium and reduced chloroalkane and chloroalkene degradation predicted high levels of FLC λ (P < 0.05).
Conclusions: Gut dysbiosis in CKD patients was confirmed by controlling for confounders in the present study. Additionally, the association between gut dysbiosis and FLC λ levels demonstrates the existence of crosstalk between the microbiome and immune response in CKD.
Keywords: Bifidobacterium; chronic kidney disease; confounders; free immunoglobulin light chains; gut microbiome.
Copyright © 2021 Liu, Xu, Chao, Chen, Shao, Sun, Hong, Hu, Jiang, Zhang, Xiao, Yan and Feng.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
Gut Mycobiome in Patients With Chronic Kidney Disease Was Altered and Associated With Immunological Profiles.Front Immunol. 2022 Jun 16;13:843695. doi: 10.3389/fimmu.2022.843695. eCollection 2022. Front Immunol. 2022. PMID: 35784313 Free PMC article.
-
Disorder of gut amino acids metabolism during CKD progression is related with gut microbiota dysbiosis and metagenome change.J Pharm Biomed Anal. 2018 Feb 5;149:425-435. doi: 10.1016/j.jpba.2017.11.040. Epub 2017 Nov 15. J Pharm Biomed Anal. 2018. PMID: 29169110
-
Blood Microbiome Profile in CKD : A Pilot Study.Clin J Am Soc Nephrol. 2019 May 7;14(5):692-701. doi: 10.2215/CJN.12161018. Epub 2019 Apr 8. Clin J Am Soc Nephrol. 2019. PMID: 30962186 Free PMC article.
-
Microbiome modulation to correct uremic toxins and to preserve kidney functions.Curr Opin Nephrol Hypertens. 2020 Jan;29(1):49-56. doi: 10.1097/MNH.0000000000000565. Curr Opin Nephrol Hypertens. 2020. PMID: 31725010 Review.
-
The clinical impact of gut microbiota in chronic kidney disease.Korean J Intern Med. 2020 Nov;35(6):1305-1316. doi: 10.3904/kjim.2020.411. Epub 2020 Sep 29. Korean J Intern Med. 2020. PMID: 32872729 Free PMC article. Review.
Cited by
-
Gut Microbiota in Chronic Kidney Disease: From Composition to Modulation towards Better Outcomes-A Systematic Review.J Clin Med. 2023 Mar 1;12(5):1948. doi: 10.3390/jcm12051948. J Clin Med. 2023. PMID: 36902734 Free PMC article. Review.
-
Causal Effects of Specific Gut Microbiota on Chronic Kidney Diseases and Renal Function-A Two-Sample Mendelian Randomization Study.Nutrients. 2023 Jan 11;15(2):360. doi: 10.3390/nu15020360. Nutrients. 2023. PMID: 36678231 Free PMC article.
-
Association Between Gut Microbiota and Chronic Kidney Disease: A Two-Sample Mendelian Randomization Study in a Chinese Population.Biomedicines. 2025 Jun 6;13(6):1397. doi: 10.3390/biomedicines13061397. Biomedicines. 2025. PMID: 40564116 Free PMC article.
-
Pediatric and adult point of view on the gut-kidney axis in CKD.Pediatr Nephrol. 2025 Jul 7. doi: 10.1007/s00467-025-06780-8. Online ahead of print. Pediatr Nephrol. 2025. PMID: 40622577 Review.
-
Alteration of Skin Microbiome in CKD Patients Is Associated With Pruritus and Renal Function.Front Cell Infect Microbiol. 2022 Jun 28;12:923581. doi: 10.3389/fcimb.2022.923581. eCollection 2022. Front Cell Infect Microbiol. 2022. PMID: 35837475 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

