Autophagy-Related Long Non-coding RNA Is a Prognostic Indicator for Bladder Cancer
- PMID: 33869042
- PMCID: PMC8049181
- DOI: 10.3389/fonc.2021.647236
Autophagy-Related Long Non-coding RNA Is a Prognostic Indicator for Bladder Cancer
Abstract
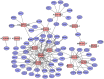
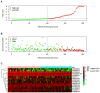
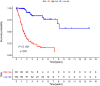
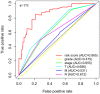
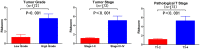
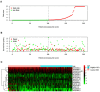
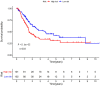
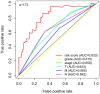
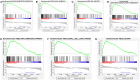
Bladder cancer (BC) is one of the most common malignant urinary system tumors, and its prognosis is poor. In recent years, autophagy has been closely linked to the development of BC. Therefore, we investigated the potential prognostic role of autophagy-related long non-coding RNA (lncRNA) in patients with BC. We obtained the lncRNA information and autophagy genes, respectively, from The Cancer Genome Atlas (TCGA) data set and the human autophagy database (HADb) and performed a co-expression analysis to identify autophagy gene-associated lncRNAs. Then, we divided the data into training group and testing group. In the training group, 15 autophagy-related lncRNAs were found to have a prognostic value (AC026369.3, USP30-as1, AC007991.2, AC104785.1, AC010503.4, AC037198.1, AC010331.1, AF131215.6, AC084357.2, THUMPD3-AS1, U62317.4, MAN1B1-DTt, AC024060.1, AL662844.4, and AC005229.4). The patients were divided into low-risk group and high-risk group based on the prognostic lncRNAs. The overall survival (OS) time for the high-risk group was shorter than that for the low-risk group [risk ratio (hazard ratio, HR) = 1.08, 95% CI: 1.06-1.10; p < 0.0001]. Using our model, the defined risk value can predict the prognosis of a patient. Next, the model was assessed in the TCGA testing group to further validate these results. A total of 203 patients with BC were recruited to verify the lncRNA characteristics. We divided these patients into high-risk group and low-risk group. The results of testing data set show that the survival time of high-risk patients is shorter than that of low-risk patients. In the training group, the area under the curve (AUC) was more than 0.7, indicating a high level of accuracy. The AUC for a risk model was greater than that for each clinical feature alone, indicating that the risk value of a model was the best indicator for predicting the prognosis. Further training data analysis showed that the gene set was significantly enriched in cancer-related pathways, including actin cytoskeleton regulation and gap junctions. In conclusion, our 15 autophagy-related lncRNAs have a prognostic potential for BC, and may play key roles in the biology of BC.
Keywords: TCGA; autophagy; bladder cancer; lncRNA; prognostic indicator.
Copyright © 2021 Wan, Guo, Fang, Xu, Hu and Luo.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
Similar articles
-
Identification and validation of an autophagy-related long non-coding RNA signature as a prognostic biomarker for patients with lung adenocarcinoma.J Thorac Dis. 2021 Feb;13(2):720-734. doi: 10.21037/jtd-20-2803. J Thorac Dis. 2021. PMID: 33717544 Free PMC article.
-
Autophagy-related long non-coding RNA prognostic model predicts prognosis and survival of melanoma patients.World J Clin Cases. 2022 Apr 16;10(11):3334-3351. doi: 10.12998/wjcc.v10.i11.3334. World J Clin Cases. 2022. PMID: 35611195 Free PMC article.
-
An autophagy-related long non-coding RNA prognostic signature accurately predicts survival outcomes in bladder urothelial carcinoma patients.Aging (Albany NY). 2020 Aug 29;12(15):15624-15637. doi: 10.18632/aging.103718. Aging (Albany NY). 2020. PMID: 32805727 Free PMC article.
-
An autophagy-related long non-coding RNA signature for glioma.FEBS Open Bio. 2019 Mar 5;9(4):653-667. doi: 10.1002/2211-5463.12601. eCollection 2019 Apr. FEBS Open Bio. 2019. PMID: 30984540 Free PMC article.
-
Critical roles of lncRNA-mediated autophagy in urologic malignancies.Front Pharmacol. 2024 Jun 13;15:1405199. doi: 10.3389/fphar.2024.1405199. eCollection 2024. Front Pharmacol. 2024. PMID: 38939836 Free PMC article. Review.
Cited by
-
Identification of the three subtypes and the prognostic characteristics of stomach adenocarcinoma: analysis of the hypoxia-related long non-coding RNAs.Funct Integr Genomics. 2022 Oct;22(5):919-936. doi: 10.1007/s10142-022-00867-3. Epub 2022 Jun 4. Funct Integr Genomics. 2022. PMID: 35665866
-
Necroptosis-related lncRNA signatures determine prognosis in breast cancer patients.Sci Rep. 2022 Jul 4;12(1):11268. doi: 10.1038/s41598-022-15209-3. Sci Rep. 2022. PMID: 35787661 Free PMC article.
-
A prognostic model based on immune-related long noncoding RNAs for patients with epithelial ovarian cancer.J Ovarian Res. 2022 Jan 15;15(1):8. doi: 10.1186/s13048-021-00930-w. J Ovarian Res. 2022. PMID: 35031063 Free PMC article.
-
T2DB: A Web Database for Long Non-Coding RNA Genes in Type II Diabetes.Noncoding RNA. 2023 May 8;9(3):30. doi: 10.3390/ncrna9030030. Noncoding RNA. 2023. PMID: 37218990 Free PMC article.
-
Construction of a Novel Lung Adenocarcinoma Immune-Related lncRNA Pair Signature.Int J Gen Med. 2021 Aug 11;14:4279-4289. doi: 10.2147/IJGM.S325240. eCollection 2021. Int J Gen Med. 2021. PMID: 34421308 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

