Role of NKG2a/c+CD8+ T cells in pathogenic versus non-pathogenic SIV infections
- PMID: 33870131
- PMCID: PMC8040270
- DOI: 10.1016/j.isci.2021.102314
Role of NKG2a/c+CD8+ T cells in pathogenic versus non-pathogenic SIV infections
Abstract
Some viruses have established an equilibrium with their host. African green monkeys (AGM) display persistent high viral replication in the blood and intestine during Simian immunodeficiency virus (SIV) infection but resolve systemic inflammation after acute infection and lack intestinal immune or tissue damage during chronic infection. We show that NKG2a/c +CD8+ T cells increase in the blood and intestine of AGM in response to SIVagm infection in contrast to SIVmac infection in macaques, the latter modeling HIV infection. NKG2a/c +CD8+ T cells were not expanded in lymph nodes, and CXCR5+NKG2a/c +CD8+ T cell frequencies further decreased after SIV infection. Genome-wide transcriptome analysis of NKG2a/c +CD8+ T cells from AGM revealed the expression of NK cell receptors, and of molecules with cytotoxic effector, gut homing, and immunoregulatory and gut barrier function, including CD73. NKG2a/c +CD8+ T cells correlated negatively with IL-23 in the intestine during SIVmac infection. The data suggest a potential regulatory role of NKG2a/c +CD8+ T cells in intestinal inflammation during SIV/HIV infections.
Keywords: Immunology; Transcriptomics; Viral Microbiology.
© 2021 The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



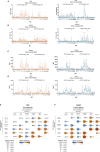



References
-
- Alter G., Malenfant J.M., Altfeld M. CD107a as a functional marker for the identification of natural killer cell activity. J. Immunol. Methods. 2004;294:15–22. - PubMed
-
- Bertone S., Schiavetti F., Bellomo R., Vitale C., Ponte M., Moretta L., Mingari M.C. Transforming growth factor-β-induced expression of CD94/NKG2A inhibitory receptors in human T lymphocytes. Eur. J. Immunol. 1999;29:23–29. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

