Structural basis for backtracking by the SARS-CoV-2 replication-transcription complex
- PMID: 33883267
- PMCID: PMC8126829
- DOI: 10.1073/pnas.2102516118
Structural basis for backtracking by the SARS-CoV-2 replication-transcription complex
Abstract
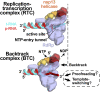
Backtracking, the reverse motion of the transcriptase enzyme on the nucleic acid template, is a universal regulatory feature of transcription in cellular organisms but its role in viruses is not established. Here we present evidence that backtracking extends into the viral realm, where backtracking by the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) RNA-dependent RNA polymerase (RdRp) may aid viral transcription and replication. Structures of SARS-CoV-2 RdRp bound to the essential nsp13 helicase and RNA suggested the helicase facilitates backtracking. We use cryo-electron microscopy, RNA-protein cross-linking, and unbiased molecular dynamics simulations to characterize SARS-CoV-2 RdRp backtracking. The results establish that the single-stranded 3' segment of the product RNA generated by backtracking extrudes through the RdRp nucleoside triphosphate (NTP) entry tunnel, that a mismatched nucleotide at the product RNA 3' end frays and enters the NTP entry tunnel to initiate backtracking, and that nsp13 stimulates RdRp backtracking. Backtracking may aid proofreading, a crucial process for SARS-CoV-2 resistance against antivirals.
Keywords: RNA-dependent RNA polymerase; backtracking; coronavirus; cryo-electron microscopy; molecular dynamics.
Copyright © 2021 the Author(s). Published by PNAS.
Conflict of interest statement
The authors declare no competing interest.
Figures





Update of
-
Structural basis for backtracking by the SARS-CoV-2 replication-transcription complex.bioRxiv [Preprint]. 2021 Mar 14:2021.03.13.435256. doi: 10.1101/2021.03.13.435256. bioRxiv. 2021. Update in: Cell. 2020 Sep 17;182(6):1560-1573.e13. doi: 10.1016/j.cell.2020.07.033. Update in: Proc Natl Acad Sci U S A. 2021 May 11;118(19):e2102516118. doi: 10.1073/pnas.2102516118. PMID: 33758867 Free PMC article. Updated. Preprint.
References
-
- US FDA , Remdesivir Emergency Use Authorization letter (2020). https://www.fda.gov/media/137564/download. Accessed 18 April 2021.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

