Histone dynamics mediate DNA unwrapping and sliding in nucleosomes
- PMID: 33888707
- PMCID: PMC8062685
- DOI: 10.1038/s41467-021-22636-9
Histone dynamics mediate DNA unwrapping and sliding in nucleosomes
Abstract
Nucleosomes are elementary building blocks of chromatin in eukaryotes. They tightly wrap ∼147 DNA base pairs around an octamer of histone proteins. How nucleosome structural dynamics affect genome functioning is not completely clear. Here we report all-atom molecular dynamics simulations of nucleosome core particles at a timescale of 15 microseconds. At this timescale, functional modes of nucleosome dynamics such as spontaneous nucleosomal DNA breathing, unwrapping, twisting, and sliding were observed. We identified atomistic mechanisms of these processes by analyzing the accompanying structural rearrangements of the histone octamer and histone-DNA contacts. Octamer dynamics and plasticity were found to enable DNA unwrapping and sliding. Through multi-scale modeling, we showed that nucleosomal DNA dynamics contribute to significant conformational variability of the chromatin fiber at the supranucleosomal level. Our study further supports mechanistic coupling between fine details of histone dynamics and chromatin functioning, provides a framework for understanding the effects of various chromatin modifications.
Conflict of interest statement
The authors declare no competing interests.
Figures


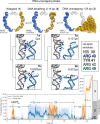






References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

