Identification of the Key Regulators of Spina Bifida Through Graph-Theoretical Approach
- PMID: 33889172
- PMCID: PMC8056047
- DOI: 10.3389/fgene.2021.597983
Identification of the Key Regulators of Spina Bifida Through Graph-Theoretical Approach
Abstract
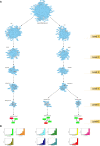
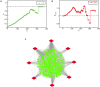
Spina Bifida (SB) is a congenital spinal cord malformation. Efforts to discern the key regulators (KRs) of the SB protein-protein interaction (PPI) network are requisite for developing its successful interventions. The architecture of the SB network, constructed from 117 manually curated genes was found to self-organize into a scale-free fractal state having a weak hierarchical organization. We identified three modules/motifs consisting of ten KRs, namely, TNIP1, TNF, TRAF1, TNRC6B, KMT2C, KMT2D, NCOA3, TRDMT1, DICER1, and HDAC1. These KRs serve as the backbone of the network, they propagate signals through the different hierarchical levels of the network to conserve the network's stability while maintaining low popularity in the network. We also observed that the SB network exhibits a rich-club organization, the formation of which is attributed to our key regulators also except for TNIP1 and TRDMT1. The KRs that were found to ally with each other and emerge in the same motif, open up a new dimension of research of studying these KRs together. Owing to the multiple etiology and mechanisms of SB, a combination of several biomarkers is expected to have higher diagnostic accuracy for SB as compared to using a single biomarker. So, if all the KRs present in a single module/motif are targetted together, they can serve as biomarkers for the diagnosis of SB. Our study puts forward some novel SB-related genes that need further experimental validation to be considered as reliable future biomarkers and therapeutic targets.
Keywords: Spina bifida; key regulators; protein-protein interaction network; rich-club analysis; topological analysis.
Copyright © 2021 Tamkeen, AlOmar, Alqahtani, Al-jurayyan, Farooqui, Tazyeen, Ahmad and Ishrat.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures






Similar articles
-
Identification of common candidate genes and pathways for Spina Bifida and Wilm's Tumor using an integrative bioinformatics analysis.J Biomol Struct Dyn. 2024 Jan-Feb;42(2):977-992. doi: 10.1080/07391102.2023.2199080. Epub 2023 Apr 13. J Biomol Struct Dyn. 2024. PMID: 37051780
-
Methodology of predicting novel key regulators in ovarian cancer network: a network theoretical approach.BMC Cancer. 2019 Nov 21;19(1):1129. doi: 10.1186/s12885-019-6309-6. BMC Cancer. 2019. PMID: 31752757 Free PMC article.
-
Transition Readiness Assessment Questionnaire Spina Bifida (TRAQ-SB) specific module and its association with clinical outcomes among youth and young adults with spina bifida.J Pediatr Rehabil Med. 2019;12(4):405-413. doi: 10.3233/PRM-180595. J Pediatr Rehabil Med. 2019. PMID: 31744032
-
[Spina bifida].Radiologe. 2018 Jul;58(7):659-663. doi: 10.1007/s00117-018-0401-9. Radiologe. 2018. PMID: 29797041 Review. German.
-
Prevalence rates of spina bifida in Alberta, Canada: 2001-2015. Can we achieve more prevention?Birth Defects Res. 2019 Feb 1;111(3):151-158. doi: 10.1002/bdr2.1438. Epub 2018 Dec 18. Birth Defects Res. 2019. PMID: 30561844 Review.
Cited by
-
Comprehensive mapping of the AOP-Wiki database: identifying biological and disease gaps.Front Toxicol. 2024 Mar 8;6:1285768. doi: 10.3389/ftox.2024.1285768. eCollection 2024. Front Toxicol. 2024. PMID: 38523647 Free PMC article.
References
-
- Albert R., Barabási A.-L. (2002). Statistical mechanics of complex networks. Rev. Mod. Phys. 74 47–97. 10.1103/RevModPhys.74.47 - DOI
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

