G2S: A New Deep Learning Tool for Predicting Stool Microbiome Structure From Oral Microbiome Data
- PMID: 33897763
- PMCID: PMC8062976
- DOI: 10.3389/fgene.2021.644516
G2S: A New Deep Learning Tool for Predicting Stool Microbiome Structure From Oral Microbiome Data
Abstract
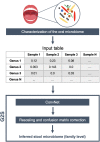
Deep learning methodologies have revolutionized prediction in many fields and show the potential to do the same in microbial metagenomics. However, deep learning is still unexplored in the field of microbiology, with only a few software designed to work with microbiome data. Within the meta-community theory, we foresee new perspectives for the development and application of deep learning algorithms in the field of the human microbiome. In this context, we developed G2S, a bioinformatic tool for taxonomic prediction of the human fecal microbiome directly from the oral microbiome data of the same individual. The tool uses a deep convolutional neural network trained on paired oral and fecal samples from populations across the globe, which allows inferring the stool microbiome at the family level more accurately than other available approaches. The tool can be used in retrospective studies, where fecal sampling was not performed, and especially in the field of paleomicrobiology, as a unique opportunity to recover data related to ancient gut microbiome configurations. G2S was validated on already characterized oral and fecal sample pairs, and then applied to ancient microbiome data from dental calculi, to derive putative intestinal components in medieval subjects.
Keywords: deep learning; gut microbiome; microbiome; oral microbiome; paleomicrobiology.
Copyright © 2021 Rampelli, Fabbrini, Candela, Biagi, Brigidi and Turroni.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


References
-
- Bishop C. M. (2016). Pattern Recognition and Machine Learning. New York, NY: Springer.
LinkOut - more resources
Full Text Sources
Other Literature Sources

