Single-cell characterization of a model of poly I:C-stimulated peripheral blood mononuclear cells in severe asthma
- PMID: 33902571
- PMCID: PMC8074196
- DOI: 10.1186/s12931-021-01709-9
Single-cell characterization of a model of poly I:C-stimulated peripheral blood mononuclear cells in severe asthma
Abstract
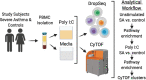
Background: Asthma has been associated with impaired interferon response. Multiple cell types have been implicated in such response impairment and may be responsible for asthma immunopathology. However, existing models to study the immune response in asthma are limited by bulk profiling of cells. Our objective was to Characterize a model of peripheral blood mononuclear cells (PBMCs) of patients with severe asthma (SA) and its response to the TLR3 agonist Poly I:C using two single-cell methods.
Methods: Two complementary single-cell methods, DropSeq for single-cell RNA sequencing (scRNA-Seq) and mass cytometry (CyTOF), were used to profile PBMCs of SA patients and healthy controls (HC). Poly I:C-stimulated and unstimulated cells were analyzed in this study.
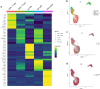
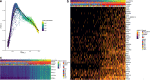
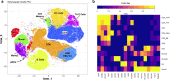
Results: PBMCs (n = 9414) from five SA (n = 6099) and three HC (n = 3315) were profiled using scRNA-Seq. Six main cell subsets, namely CD4 + T cells, CD8 + T cells, natural killer (NK) cells, B cells, dendritic cells (DCs), and monocytes, were identified. CD4 + T cells were the main cell type in SA and demonstrated a pro-inflammatory profile characterized by increased JAK1 expression. Following Poly I:C stimulation, PBMCs from SA had a robust induction of interferon pathways compared with HC. CyTOF profiling of Poly I:C stimulated and unstimulated PBMCs (n = 160,000) from the same individuals (SA = 5; HC = 3) demonstrated higher CD8 + and CD8 + effector T cells in SA at baseline, followed by a decrease of CD8 + effector T cells after poly I:C stimulation.
Conclusions: Single-cell profiling of an in vitro model using PBMCs in patients with SA identified activation of pro-inflammatory pathways at baseline and strong response to Poly I:C, as well as quantitative changes in CD8 + effector cells. Thus, transcriptomic and cell quantitative changes are associated with immune cell heterogeneity in this model to evaluate interferon responses in severe asthma.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures


Similar articles
-
Single-cell RNA sequencing reveals transcriptional changes in circulating immune cells from patients with severe asthma induced by biologics.Exp Mol Med. 2024 Dec;56(12):2755-2762. doi: 10.1038/s12276-024-01368-y. Epub 2024 Dec 13. Exp Mol Med. 2024. PMID: 39672815 Free PMC article.
-
A single-cell atlas of the peripheral immune response in patients with severe COVID-19.Nat Med. 2020 Jul;26(7):1070-1076. doi: 10.1038/s41591-020-0944-y. Epub 2020 Jun 8. Nat Med. 2020. PMID: 32514174 Free PMC article.
-
A distinct immune landscape in anti-synthetase syndrome profiled by a single-cell genomic study.Front Immunol. 2024 Oct 24;15:1436114. doi: 10.3389/fimmu.2024.1436114. eCollection 2024. Front Immunol. 2024. PMID: 39512337 Free PMC article.
-
Heterogeneity of immune cells in human atherosclerosis revealed by scRNA-Seq.Cardiovasc Res. 2021 Nov 22;117(13):2537-2543. doi: 10.1093/cvr/cvab260. Cardiovasc Res. 2021. PMID: 34343272 Free PMC article. Review.
-
Single-Cell Analysis: A Method for In-Depth Phenotyping of Cells Involved in Asthma.Int J Mol Sci. 2024 Nov 25;25(23):12633. doi: 10.3390/ijms252312633. Int J Mol Sci. 2024. PMID: 39684345 Free PMC article. Review.
Cited by
-
Immunostimulant Bathing Influences the Expression of Immune- and Metabolic-Related Genes in Atlantic Salmon Alevins.Biology (Basel). 2021 Sep 29;10(10):980. doi: 10.3390/biology10100980. Biology (Basel). 2021. PMID: 34681079 Free PMC article.
-
Functional immunophenotyping of children with critical status asthmaticus identifies differential gene expression responses in neutrophils exposed to a poly(I:C) stimulus.Sci Rep. 2022 Nov 16;12(1):19644. doi: 10.1038/s41598-022-24261-y. Sci Rep. 2022. PMID: 36385161 Free PMC article.
-
Single-cell RNA sequencing reveals transcriptional changes in circulating immune cells from patients with severe asthma induced by biologics.Exp Mol Med. 2024 Dec;56(12):2755-2762. doi: 10.1038/s12276-024-01368-y. Epub 2024 Dec 13. Exp Mol Med. 2024. PMID: 39672815 Free PMC article.
-
Single-cell RNA-sequencing in asthma research.Front Immunol. 2022 Nov 29;13:988573. doi: 10.3389/fimmu.2022.988573. eCollection 2022. Front Immunol. 2022. PMID: 36524132 Free PMC article. Review.
-
Identification of potential biomarkers and immune infiltration characteristics in severe asthma.Int J Immunopathol Pharmacol. 2022 Jan-Dec;36:3946320221114194. doi: 10.1177/03946320221114194. Int J Immunopathol Pharmacol. 2022. PMID: 35817495 Free PMC article.
References
MeSH terms
Substances
Grants and funding
- R21 LM012884/LM/NLM NIH HHS/United States
- 113393/Flight Attendant Medical Research Institute
- R01 HL141237/HL/NHLBI NIH HHS/United States
- UL1 TR001863/TR/NCATS NIH HHS/United States
- K01HL125474/HL/NHLBI NIH HHS/United States
- R21 LM012884/NH/NIH HHS/United States
- R56 HL141237/HL/NHLBI NIH HHS/United States
- K08 HL135402/HL/NHLBI NIH HHS/United States
- R01 HL118346/NH/NIH HHS/United States
- K01 HL125514/HL/NHLBI NIH HHS/United States
- R01 HL153604/HL/NHLBI NIH HHS/United States
- R03 HL154275/HL/NHLBI NIH HHS/United States
- K01 HL125474/HL/NHLBI NIH HHS/United States
- R01 HL126094/NH/NIH HHS/United States
- R01 HL141237/NH/NIH HHS/United States
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

