Learning fuzzy clustering for SPECT/CT segmentation via convolutional neural networks
- PMID: 33905560
- PMCID: PMC9973404
- DOI: 10.1002/mp.14903
Learning fuzzy clustering for SPECT/CT segmentation via convolutional neural networks
Abstract
Purpose: Quantitative bone single-photon emission computed tomography (QBSPECT) has the potential to provide a better quantitative assessment of bone metastasis than planar bone scintigraphy due to its ability to better quantify activity in overlapping structures. An important element of assessing the response of bone metastasis is accurate image segmentation. However, limited by the properties of QBSPECT images, the segmentation of anatomical regions-of-interests (ROIs) still relies heavily on the manual delineation by experts. This work proposes a fast and robust automated segmentation method for partitioning a QBSPECT image into lesion, bone, and background.
Methods: We present a new unsupervised segmentation loss function and its semi- and supervised variants for training a convolutional neural network (ConvNet). The loss functions were developed based on the objective function of the classical Fuzzy C-means (FCM) algorithm. The first proposed loss function can be computed within the input image itself without any ground truth labels, and is thus unsupervised; the proposed supervised loss function follows the traditional paradigm of the deep learning-based segmentation methods and leverages ground truth labels during training. The last loss function is a combination of the first and the second and includes a weighting parameter, which enables semi-supervised segmentation using deep learning neural network.
Experiments and results: We conducted a comprehensive study to compare our proposed methods with ConvNets trained using supervised, cross-entropy and Dice loss functions, and conventional clustering methods. The Dice similarity coefficient (DSC) and several other metrics were used as figures of merit as applied to the task of delineating lesion and bone in both simulated and clinical SPECT/CT images. We experimentally demonstrated that the proposed methods yielded good segmentation results on a clinical dataset even though the training was done using realistic simulated images. On simulated SPECT/CT, the proposed unsupervised model's accuracy was greater than the conventional clustering methods while reducing computation time by 200-fold. For the clinical QBSPECT/CT, the proposed semi-supervised ConvNet model, trained using simulated images, produced DSCs of and for lesion and bone segmentation in SPECT, and a DSC of bone segmentation of CT images. These DSCs were larger than that for standard segmentation loss functions by for SPECT segmentation, and for CT segmentation with P-values from a paired t-test.
Conclusions: A ConvNet-based image segmentation method that uses novel loss functions was developed and evaluated. The method can operate in unsupervised, semi-supervised, or fully-supervised modes depending on the availability of annotated training data. The results demonstrated that the proposed method provides fast and robust lesion and bone segmentation for QBSPECT/CT. The method can potentially be applied to other medical image segmentation applications.
Keywords: convolutional neural networks; fuzzy C-means; image segmentation; nuclear medicine.
© 2021 The Authors. Medical Physics published by Wiley Periodicals LLC on behalf of American Association of Physicists in Medicine.
Figures
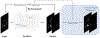
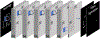
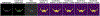


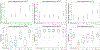




Similar articles
-
Semi-supervised abdominal multi-organ segmentation by object-redrawing.Med Phys. 2024 Nov;51(11):8334-8347. doi: 10.1002/mp.17364. Epub 2024 Aug 21. Med Phys. 2024. PMID: 39167059
-
Automated segmentation of lesions and organs at risk on [68Ga]Ga-PSMA-11 PET/CT images using self-supervised learning with Swin UNETR.Cancer Imaging. 2024 Feb 29;24(1):30. doi: 10.1186/s40644-024-00675-x. Cancer Imaging. 2024. PMID: 38424612 Free PMC article.
-
Lung tumor segmentation in 4D CT images using motion convolutional neural networks.Med Phys. 2021 Nov;48(11):7141-7153. doi: 10.1002/mp.15204. Epub 2021 Sep 13. Med Phys. 2021. PMID: 34469001 Free PMC article.
-
Automated vessel segmentation in lung CT and CTA images via deep neural networks.J Xray Sci Technol. 2021;29(6):1123-1137. doi: 10.3233/XST-210955. J Xray Sci Technol. 2021. PMID: 34421004 Review.
-
A deep learning-based approach to automatic proximal femur segmentation in quantitative CT images.Med Biol Eng Comput. 2022 May;60(5):1417-1429. doi: 10.1007/s11517-022-02529-9. Epub 2022 Mar 24. Med Biol Eng Comput. 2022. PMID: 35322343 Review.
Cited by
-
Remote sensing image information extraction based on Compensated Fuzzy Neural Network and big data analytics.BMC Med Imaging. 2024 Apr 10;24(1):86. doi: 10.1186/s12880-024-01266-9. BMC Med Imaging. 2024. PMID: 38600525 Free PMC article.
-
SwinCross: Cross-modal Swin transformer for head-and-neck tumor segmentation in PET/CT images.Med Phys. 2024 Mar;51(3):2096-2107. doi: 10.1002/mp.16703. Epub 2023 Sep 30. Med Phys. 2024. PMID: 37776263 Free PMC article.
-
Semi-supervised learning towards automated segmentation of PET images with limited annotations: application to lymphoma patients.Phys Eng Sci Med. 2024 Sep;47(3):833-849. doi: 10.1007/s13246-024-01408-x. Epub 2024 Mar 21. Phys Eng Sci Med. 2024. PMID: 38512435
-
Pix2Pix generative adversarial network for low dose myocardial perfusion SPECT denoising.Quant Imaging Med Surg. 2022 Jul;12(7):3539-3555. doi: 10.21037/qims-21-1042. Quant Imaging Med Surg. 2022. PMID: 35782241 Free PMC article.
-
Computed Tomography Images under Artificial Intelligence Algorithms on the Treatment Evaluation of Intracerebral Hemorrhage with Minimally Invasive Aspiration.Comput Math Methods Med. 2022 Apr 22;2022:6204089. doi: 10.1155/2022/6204089. eCollection 2022. Comput Math Methods Med. 2022. Retraction in: Comput Math Methods Med. 2023 Dec 13;2023:9802621. doi: 10.1155/2023/9802621. PMID: 35495888 Free PMC article. Retracted.
References
-
- Siegel RL, Miller KD, and Jemal A, “Cancer statistics, 2020,” CA: A Cancer Journal for Clinicians, vol. 70, no. 1, pp. 7–30, jan 2020. [Online]. Available: https://onlinelibrary.wiley.com/doi/abs/10.3322/caac.21590 - DOI - PubMed
-
- He B et al., “Evaluation of quantitative imaging methods for organ activity and residence time estimation using a population of phantoms having realistic variations in anatomy and uptake,” Medical Physics, vol. 36, no. 2, pp. 612–619, 2009. [Online]. Available: https://aapm.onlinelibrary.wiley.com/doi/abs/10.1118/1.3063156 - DOI - PMC - PubMed
-
- He B and Frey EC, “Comparison of conventional, model-based quantitative planar, and quantitative SPECT image processing methods for organ activity estimation using In-111 agents,” Physics in Medicine and Biology, vol. 51, no. 16, pp. 3967–3981, aug 2006. [Online]. Available: https://iopscience.iop.org/article/10.1088/0031-9155/51/16/006 - DOI - PubMed
-
- He B, Du Y, Song X, Segars WP, and Frey EC, “A Monte Carlo and physical phantom evaluation of quantitative In-111 SPECT,” Physics in Medicine and Biology, vol. 50, no. 17, pp. 4169–4185, sep 2005. - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

