Synergistic DNA- and Protein-Based Recognition Promote an RNA-Templated Bio-orthogonal Reaction
- PMID: 33914384
- PMCID: PMC8362040
- DOI: 10.1002/chem.202101103
Synergistic DNA- and Protein-Based Recognition Promote an RNA-Templated Bio-orthogonal Reaction
Abstract
Biomolecular assemblies composed of proteins and oligonucleotides play a central role in biological processes. While in nature, oligonucleotides and proteins usually assemble via non-covalent interactions, synthetic conjugates have been developed which covalently link both modalities. The resulting peptide-oligonucleotide conjugates have facilitated novel biological applications as well as the design of functional supramolecular systems and materials. However, despite the importance of concerted protein/oligonucleotide recognition in nature, conjugation approaches have barely utilized the synergistic recognition abilities of such complexes. Herein, the structure-based design of peptide-DNA conjugates that bind RNA through Watson-Crick base pairing combined with peptide-mediated major groove recognition is reported. Two distinct conjugate families with tunable binding characteristics have been designed to adjacently bind a particular RNA sequence. In the resulting ternary complex, their peptide elements are located in proximity, a feature that was used to enable an RNA-templated click reaction. The introduced structure-based design approach opens the door to novel functional biomolecular assemblies.
Keywords: RNA recognition; bio-conjugation; peptides; self-assembly; strain-promoted click reaction.
© 2021 The Authors. Chemistry - A European Journal published by Wiley-VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



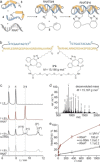
References
-
- J. Wilce, J. Vivian, M. Wilce, in Protein Dimerization and Oligomerization in Biology (Ed. J. M. Matthews), Springer, New York, 2012, pp. 91–104.
-
- Zhao D., Kong Y., Zhao S., Xing H., Top. Curr. Chem. 2020, 378, 41. - PubMed
-
- Dovgan I., Koniev O., Kolodych S., Wagner A., Bioconjugate Chem. 2019, 30, 2483–2501. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

