Genetic Diversity of SARS-CoV-2 over a One-Year Period of the COVID-19 Pandemic: A Global Perspective
- PMID: 33920487
- PMCID: PMC8069977
- DOI: 10.3390/biomedicines9040412
Genetic Diversity of SARS-CoV-2 over a One-Year Period of the COVID-19 Pandemic: A Global Perspective
Abstract
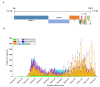
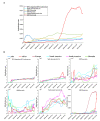
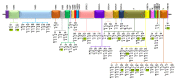
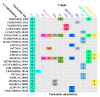
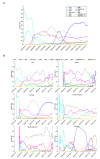
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) caused a global pandemic of coronavirus disease in 2019 (COVID-19). Genome surveillance is a key method to track the spread of SARS-CoV-2 variants. Genetic diversity and evolution of SARS-CoV-2 were analyzed based on 260,673 whole-genome sequences, which were sampled from 62 countries between 24 December 2019 and 12 January 2021. We found that amino acid (AA) substitutions were observed in all SARS-CoV-2 proteins, and the top six proteins with the highest substitution rates were ORF10, nucleocapsid, ORF3a, spike glycoprotein, RNA-dependent RNA polymerase, and ORF8. Among 25,629 amino acid substitutions at 8484 polymorphic sites across the coding region of the SARS-CoV-2 genome, the D614G (93.88%) variant in spike and the P323L (93.74%) variant in RNA-dependent RNA polymerase were the dominant variants on six continents. As of January 2021, the genomic sequences of SARS-CoV-2 could be divided into at least 12 different clades. Distributions of SARS-CoV-2 clades were featured with temporal and geographical dynamics on six continents. Overall, this large-scale analysis provides a detailed mapping of SARS-CoV-2 variants in different geographic areas at different time points, highlighting the importance of evaluating highly prevalent variants in the development of SARS-CoV-2 antiviral drugs and vaccines.
Keywords: COVID-19; SARS-CoV-2; genetic diversity; genetic variant; global pandemic.
Conflict of interest statement
None of Erik De Clercq’s drugs [70,71] were designed or approved for coronavirus treatment. The authors declare no conflict of interests.
Figures

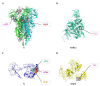
Similar articles
-
Global variation in SARS-CoV-2 proteome and its implication in pre-lockdown emergence and dissemination of 5 dominant SARS-CoV-2 clades.Infect Genet Evol. 2021 Sep;93:104973. doi: 10.1016/j.meegid.2021.104973. Epub 2021 Jun 18. Infect Genet Evol. 2021. PMID: 34147651 Free PMC article.
-
The extent of molecular variation in novel SARS-CoV-2 after the six-month global spread.Infect Genet Evol. 2021 Jul;91:104800. doi: 10.1016/j.meegid.2021.104800. Epub 2021 Mar 5. Infect Genet Evol. 2021. PMID: 33677109 Free PMC article.
-
Contrasting Epidemiology and Population Genetics of COVID-19 Infections Defined by Multilocus Genotypes in SARS-CoV-2 Genomes Sampled Globally.Viruses. 2022 Jun 29;14(7):1434. doi: 10.3390/v14071434. Viruses. 2022. PMID: 35891414 Free PMC article.
-
Emergence, evolution, and vaccine production approaches of SARS-CoV-2 virus: Benefits of getting vaccinated and common questions.Saudi J Biol Sci. 2022 Apr;29(4):1981-1997. doi: 10.1016/j.sjbs.2021.12.020. Epub 2021 Dec 13. Saudi J Biol Sci. 2022. PMID: 34924802 Free PMC article. Review.
-
Characterization of SARS-CoV-2 different variants and related morbidity and mortality: a systematic review.Eur J Med Res. 2021 Jun 8;26(1):51. doi: 10.1186/s40001-021-00524-8. Eur J Med Res. 2021. PMID: 34103090 Free PMC article.
Cited by
-
Phylogenetic and genome-wide mutational analysis of SARS-CoV-2 strains circulating in Nigeria: no implications for attenuated COVID-19 outcomes.Osong Public Health Res Perspect. 2022 Apr;13(2):101-113. doi: 10.24171/j.phrp.2021.0329. Epub 2022 Apr 22. Osong Public Health Res Perspect. 2022. PMID: 35538682 Free PMC article.
-
IgG-like bispecific antibodies with potent and synergistic neutralization against circulating SARS-CoV-2 variants of concern.Nat Commun. 2022 Oct 3;13(1):5814. doi: 10.1038/s41467-022-33030-4. Nat Commun. 2022. PMID: 36192374 Free PMC article.
-
Association between inflammatory cytokines and anti-SARS-CoV-2 antibodies in hospitalized patients with COVID-19.Immun Ageing. 2022 Mar 5;19(1):12. doi: 10.1186/s12979-022-00271-2. Immun Ageing. 2022. PMID: 35248063 Free PMC article.
-
COVID-19 in China and the US: Differences in Hospital Admission Co-Variates and Outcomes.Vaccines (Basel). 2022 Feb 18;10(2):326. doi: 10.3390/vaccines10020326. Vaccines (Basel). 2022. PMID: 35214784 Free PMC article.
-
The importance of accessory protein variants in the pathogenicity of SARS-CoV-2.Arch Biochem Biophys. 2022 Mar 15;717:109124. doi: 10.1016/j.abb.2022.109124. Epub 2022 Jan 24. Arch Biochem Biophys. 2022. PMID: 35085577 Free PMC article.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

