Eps15 Homology Domain-Containing Protein 3 Hypermethylation as a Prognostic and Predictive Marker for Colorectal Cancer
- PMID: 33922189
- PMCID: PMC8145505
- DOI: 10.3390/biomedicines9050453
Eps15 Homology Domain-Containing Protein 3 Hypermethylation as a Prognostic and Predictive Marker for Colorectal Cancer
Abstract
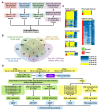
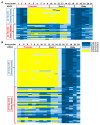
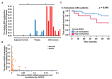
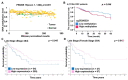
Colorectal cancer (CRC) arises from chromosomal instability, resulting from aberrant hypermethylation in tumor suppressor genes. This study identified hypermethylated genes in CRC and investigated how they affect clinical outcomes. Methylation levels of specific genes were analyzed from The Cancer Genome Atlas dataset and 20 breast cancer, 16 esophageal cancer, 33 lung cancer, 15 uterine cancer, 504 CRC, and 9 colon polyp tissues and 102 CRC plasma samples from a Taiwanese cohort. In the Asian cohort, Eps15 homology domain-containing protein 3 (EHD3) had twofold higher methylation in 44.4% of patients with colonic polyps, 37.3% of plasma from CRC patients, and 72.6% of CRC tissues, which was connected to vascular invasion and high microsatellite instability. Furthermore, EHD3 hypermethylation was detected in other gastrointestinal cancers. In the Asian CRC cohort, low EHD3 mRNA expression was found in 45.1% of patients and was connected to lymph node metastasis. Multivariate Cox proportional-hazards survival analysis revealed that hypermethylation in women and low mRNA expression were associated with overall survival. In the Western CRC cohort, EHD3 hypermethylation was also connected to overall survival and lower chemotherapy and antimetabolite response rates. In conclusion, EHD3 hypermethylation contributes to the development of CRC in both Asian and Western populations.
Keywords: DNA methylation; EHD3; circulating cell-free DNA (ccfDNA); colorectal cancer (CRC); early detection; prognostic marker.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Similar articles
-
Hypermethylation and decreased expression of TMEM240 are potential early-onset biomarkers for colorectal cancer detection, poor prognosis, and early recurrence prediction.Clin Epigenetics. 2020 May 12;12(1):67. doi: 10.1186/s13148-020-00855-z. Clin Epigenetics. 2020. PMID: 32398064 Free PMC article.
-
Hypermethylation of BEND5 contributes to cell proliferation and is a prognostic marker of colorectal cancer.Oncotarget. 2017 Nov 1;8(69):113431-113443. doi: 10.18632/oncotarget.22266. eCollection 2017 Dec 26. Oncotarget. 2017. PMID: 29371920 Free PMC article.
-
Clinical Relevance of Plasma DNA Methylation in Colorectal Cancer Patients Identified by Using a Genome-Wide High-Resolution Array.Ann Surg Oncol. 2015 Dec;22 Suppl 3:S1419-27. doi: 10.1245/s10434-014-4277-2. Epub 2014 Dec 4. Ann Surg Oncol. 2015. PMID: 25472652
-
Clinicopathological significance and potential drug target of O6-methylguanine-DNA methyltransferase in colorectal cancer: a meta-analysis.Tumour Biol. 2015 Aug;36(8):5839-48. doi: 10.1007/s13277-015-3254-0. Epub 2015 Feb 27. Tumour Biol. 2015. PMID: 25716203
-
APC hypermethylation for early diagnosis of colorectal cancer: a meta-analysis and literature review.Oncotarget. 2017 Jul 11;8(28):46468-46479. doi: 10.18632/oncotarget.17576. Oncotarget. 2017. PMID: 28515349 Free PMC article. Review.
Cited by
-
Genomic and Transcriptomic Research in the Discovery and Application of Colorectal Cancer Circulating Markers.Int J Mol Sci. 2023 Aug 3;24(15):12407. doi: 10.3390/ijms241512407. Int J Mol Sci. 2023. PMID: 37569782 Free PMC article. Review.
-
EHD3 promotes gastric cancer progression via Wnt/β-catenin/EMT pathway and associates with clinical prognosis and immune infiltration.Am J Cancer Res. 2023 Sep 15;13(9):4401-4417. eCollection 2023. Am J Cancer Res. 2023. PMID: 37818061 Free PMC article.
References
-
- Ministry of Health and Welfare . Statistical Results of Deaths of Taiwanese People in the 107th Year of the Republic Era. Ministry of Health and Welfare; Taiwan, China: 2019.
-
- Muller M., Hansmannel F., Arnone D., Choukour M., Ndiaye N.C., Kokten T., Houlgatte R., Peyrin-Biroulet L. Genomic and molecular alterations in human inflammatory bowel disease-associated colorectal cancer. United Eur. Gastroenterol. J. 2020;8:675–684. doi: 10.1177/2050640620919254. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

