The Role of Cell Tracing and Fate Mapping Experiments in Cardiac Outflow Tract Development, New Opportunities through Emerging Technologies
- PMID: 33925811
- PMCID: PMC8146276
- DOI: 10.3390/jcdd8050047
The Role of Cell Tracing and Fate Mapping Experiments in Cardiac Outflow Tract Development, New Opportunities through Emerging Technologies
Abstract
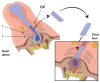
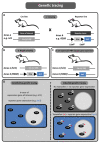
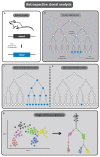
Whilst knowledge regarding the pathophysiology of congenital heart disease (CHDs) has advanced greatly in recent years, the underlying developmental processes affecting the cardiac outflow tract (OFT) such as bicuspid aortic valve, tetralogy of Fallot and transposition of the great arteries remain poorly understood. Common among CHDs affecting the OFT, is a large variation in disease phenotypes. Even though the different cell lineages contributing to OFT development have been studied for many decades, it remains challenging to relate cell lineage dynamics to the morphologic variation observed in OFT pathologies. We postulate that the variation observed in cellular contribution in these congenital heart diseases might be related to underlying cell lineage dynamics of which little is known. We believe this gap in knowledge is mainly the result of technical limitations in experimental methods used for cell lineage analysis. The aim of this review is to provide an overview of historical fate mapping and cell tracing techniques used to study OFT development and introduce emerging technologies which provide new opportunities that will aid our understanding of the cellular dynamics underlying OFT pathology.
Keywords: bicuspid aortic valve; cardiac development; cell identity; congenital heart disease; developmental biology; lineage tracing; outflow tract.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Outflow Tract Formation-Embryonic Origins of Conotruncal Congenital Heart Disease.J Cardiovasc Dev Dis. 2021 Apr 9;8(4):42. doi: 10.3390/jcdd8040042. J Cardiovasc Dev Dis. 2021. PMID: 33918884 Free PMC article. Review.
-
A Multi-Omics Approach Using a Mouse Model of Cardiac Malformations for Prioritization of Human Congenital Heart Disease Contributing Genes.Front Cardiovasc Med. 2021 Aug 24;8:683074. doi: 10.3389/fcvm.2021.683074. eCollection 2021. Front Cardiovasc Med. 2021. PMID: 34504875 Free PMC article.
-
Cardiac outflow tract anomalies.Wiley Interdiscip Rev Dev Biol. 2013 Jul;2(4):499-530. doi: 10.1002/wdev.98. Epub 2013 Feb 19. Wiley Interdiscip Rev Dev Biol. 2013. PMID: 24014420 Free PMC article. Review.
-
HAND transcription factors cooperatively specify the aorta and pulmonary trunk.Dev Biol. 2021 Aug;476:1-10. doi: 10.1016/j.ydbio.2021.03.011. Epub 2021 Mar 20. Dev Biol. 2021. PMID: 33757801 Free PMC article.
-
Nipbl Haploinsufficiency Leads to Delayed Outflow Tract Septation and Aortic Valve Thickening.Int J Mol Sci. 2023 Oct 25;24(21):15564. doi: 10.3390/ijms242115564. Int J Mol Sci. 2023. PMID: 37958548 Free PMC article.
Cited by
-
A cardioimmunologist's toolkit: genetic tools to dissect immune cells in cardiac disease.Nat Rev Cardiol. 2022 Jun;19(6):395-413. doi: 10.1038/s41569-022-00701-0. Epub 2022 May 6. Nat Rev Cardiol. 2022. PMID: 35523863 Review.
-
Introduction to Special Issue "Leaders in Cardiovascular Research, Dedicated to the Memory of Professor Adriana Gittenberger-de Groot".J Cardiovasc Dev Dis. 2022 Mar 23;9(4):92. doi: 10.3390/jcdd9040092. J Cardiovasc Dev Dis. 2022. PMID: 35448068 Free PMC article.
References
-
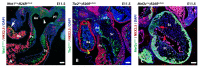
- Peterson J.C., Chughtai M., Wisse L.J., Gittenberger-de Groot A.C., Feng Q., Goumans M.-J.T.H., VanMunsteren J.C., Jongbloed M.R.M., DeRuiter M.C. Bicuspid aortic valve formation: Nos3 mutation leads to abnormal lineage patterning of neural crest cells and the second heart field. Dis. Model. Mech. 2018;11:655–658. doi: 10.1242/dmm.034637. - DOI - PMC - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

