Species-Specific Quality Control, Assembly and Contamination Detection in Microbial Isolate Sequences with AQUAMIS
- PMID: 33926025
- PMCID: PMC8145556
- DOI: 10.3390/genes12050644
Species-Specific Quality Control, Assembly and Contamination Detection in Microbial Isolate Sequences with AQUAMIS
Abstract
Sequencing of whole microbial genomes has become a standard procedure for cluster detection, source tracking, outbreak investigation and surveillance of many microorganisms. An increasing number of laboratories are currently in a transition phase from classical methods towards next generation sequencing, generating unprecedented amounts of data. Since the precision of downstream analyses depends significantly on the quality of raw data generated on the sequencing instrument, a comprehensive, meaningful primary quality control is indispensable. Here, we present AQUAMIS, a Snakemake workflow for an extensive quality control and assembly of raw Illumina sequencing data, allowing laboratories to automatize the initial analysis of their microbial whole-genome sequencing data. AQUAMIS performs all steps of primary sequence analysis, consisting of read trimming, read quality control (QC), taxonomic classification, de-novo assembly, reference identification, assembly QC and contamination detection, both on the read and assembly level. The results are visualized in an interactive HTML report including species-specific QC thresholds, allowing non-bioinformaticians to assess the quality of sequencing experiments at a glance. All results are also available as a standard-compliant JSON file, facilitating easy downstream analyses and data exchange. We have applied AQUAMIS to analyze ~13,000 microbial isolates as well as ~1000 in-silico contaminated datasets, proving the workflow's ability to perform in high throughput routine sequencing environments and reliably predict contaminations. We found that intergenus and intragenus contaminations can be detected most accurately using a combination of different QC metrics available within AQUAMIS.
Keywords: assembly; contamination; interoperability; isolate sequencing; next generation sequencing; pipeline; quality control; reproducibility; whole genome sequencing.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



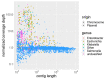
References
-
- Timme R.E., Wolfgang W.J., Balkey M., Venkata S.L.G., Randolph R., Allard M., Strain E. Optimizing open data to support one health: Best practices to ensure interoperability of genomic data from bacterial pathogens. One Health Outlook. 2020;2:20. doi: 10.1186/s42522-020-00026-3. - DOI - PMC - PubMed
-
- Bogaerts B., Nouws S., Verhaegen B., Denayer S., Van Braekel J., Winand R., Fu Q., Crombe F., Pierard D., Marchal K., et al. Validation strategy of a bioinformatics whole genome sequencing workflow for Shiga toxin-producing Escherichia coli using a reference collection extensively characterized with conventional methods. Microb. Genom. 2021 doi: 10.1099/mgen.0.000531. - DOI - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

