Estimands in epigenome-wide association studies
- PMID: 33926513
- PMCID: PMC8086103
- DOI: 10.1186/s13148-021-01083-9
Estimands in epigenome-wide association studies
Abstract
Background: In DNA methylation analyses like epigenome-wide association studies, effects in differentially methylated CpG sites are assessed. Two kinds of outcomes can be used for statistical analysis: Beta-values and M-values. M-values follow a normal distribution and help to detect differentially methylated CpG sites. As biological effect measures, differences of M-values are more or less meaningless. Beta-values are of more interest since they can be interpreted directly as differences in percentage of DNA methylation at a given CpG site, but they have poor statistical properties. Different frameworks are proposed for reporting estimands in DNA methylation analysis, relying on Beta-values, M-values, or both.
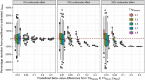
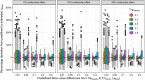
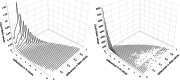
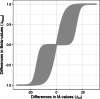
Results: We present and discuss four possible approaches of achieving estimands in DNA methylation analysis. In addition, we present the usage of M-values or Beta-values in the context of bioinformatical pipelines, which often demand a predefined outcome. We show the dependencies between the differences in M-values to differences in Beta-values in two data simulations: a analysis with and without confounder effect. Without present confounder effects, M-values can be used for the statistical analysis and Beta-values statistics for the reporting. If confounder effects exist, we demonstrate the deviations and correct the effects by the intercept method. Finally, we demonstrate the theoretical problem on two large human genome-wide DNA methylation datasets to verify the results.
Conclusions: The usage of M-values in the analysis of DNA methylation data will produce effect estimates, which cannot be biologically interpreted. The parallel usage of Beta-value statistics ignores possible confounder effects and can therefore not be recommended. Hence, if the differences in Beta-values are the focus of the study, the intercept method is recommendable. Hyper- or hypomethylated CpG sites must then be carefully evaluated. If an exploratory analysis of possible CpG sites is the aim of the study, M-values can be used for inference.
Keywords: DNA methylation; Epigenome-wide association study (EWAS); Estimands; Multiple testing; Reproducible research.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures

References
-
- Herrel A, Joly D, Danchin E. Epigenetics in ecology and evolution. Hoboken: Wiley Online Library; 2020.
-
- Heiss JA, Brennan KJ, Baccarelli AA, Téllez-Rojo MM, Estrada-Gutiérrez G, Wright RO, Just AC. Battle of epigenetic proportions: comparing illumina’s epic methylation microarrays and truseq targeted bisulfite sequencing. Epigenetics. 2020;15(1–2):174–182. doi: 10.1080/15592294.2019.1656159. - DOI - PMC - PubMed
-
- Betensky RA. The p value requires context, not a threshold. Am Stat. 2019;73(sup1):115–117. doi: 10.1080/00031305.2018.1529624. - DOI
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

