Employing core regulatory circuits to define cell identity
- PMID: 33934382
- PMCID: PMC8126924
- DOI: 10.15252/embj.2020106785
Employing core regulatory circuits to define cell identity
Abstract
The interplay between extrinsic signaling and downstream gene networks controls the establishment of cell identity during development and its maintenance in adult life. Advances in next-generation sequencing and single-cell technologies have revealed additional layers of complexity in cell identity. Here, we review our current understanding of transcription factor (TF) networks as key determinants of cell identity. We discuss the concept of the core regulatory circuit as a set of TFs and interacting factors that together define the gene expression profile of the cell. We propose the core regulatory circuit as a comprehensive conceptual framework for defining cellular identity and discuss its connections to cell function in different contexts.
Keywords: GRN; cell identity; core regulatory circuit; regenerative medicine; transcription factor.
© 2021 The Authors. Published under the terms of the CC BY 4.0 license.
Conflict of interest statement
F.M.W. is currently on secondment as executive chair of the UK Medical Research Council. The other authors declare that they have no conflict of interest.
Figures

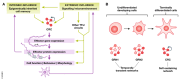

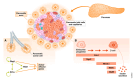
References
-
- Arendt D (2008) The evolution of cell types in animals: Emerging principles from molecular studies. Nat Rev Genet 9: 868–882 - PubMed
-
- Arendt D, Musser JM, Baker CVH, Bergman A, Cepko C, Erwin DH, Pavlicev M, Schlosser G, Widder S, Laubichler MD et al (2016) The origin and evolution of cell types. Nat Rev Genet 17: 744–757 - PubMed
-
- Arlotta P, Paşca SP (2019) Cell diversity in the human cerebral cortex: from the embryo to brain organoids. Curr Opin Neurobiol 56: 194–198 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

