Molecular Self-Assembly and Supramolecular Chemistry of Cyclic Peptides
- PMID: 33938738
- PMCID: PMC8824434
- DOI: 10.1021/acs.chemrev.0c01291
Molecular Self-Assembly and Supramolecular Chemistry of Cyclic Peptides
Abstract
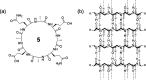
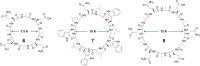
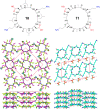
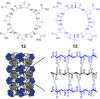
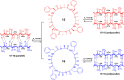
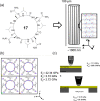
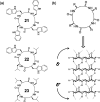
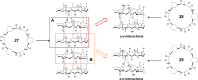
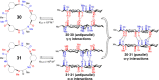
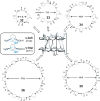
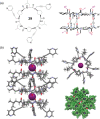
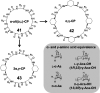
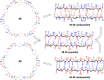
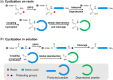
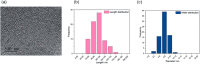
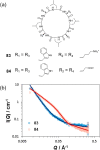
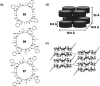
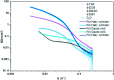
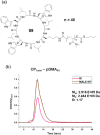
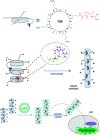
This Review focuses on the establishment and development of self-assemblies governed by the supramolecular interactions between cyclic peptides. The Review first describes the type of cyclic peptides able to assemble into tubular structures to form supramolecular cyclic peptide nanotubes. A range of cyclic peptides have been identified to have such properties, including α-peptides, β-peptides, α,γ-peptides, and peptides based on δ- and ε-amino acids. The Review covers the design and functionalization of these cyclic peptides and expands to a recent advance in the design and application of these materials through their conjugation to polymer chains to generate cyclic peptide-polymer conjugates nanostructures. The Review, then, concentrates on the challenges in characterizing these systems and presents an overview of the various analytical and characterization techniques used to date. This overview concludes with a critical survey of the various applications of the nanomaterials obtained from supramolecular cyclic peptide nanotubes, with a focus on biological and medical applications, ranging from ion channels and membrane insertion to antibacterial materials, anticancer drug delivery, gene delivery, and antiviral applications.
Conflict of interest statement
The authors declare no competing financial interest.
Figures






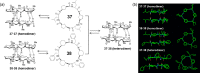




























References
-
- Lehn J.-M. Supramolecular Polymer Chemistry—Scope and Perspectives. Polym. Int. 2002, 51, 825–839. 10.1002/pi.852. - DOI
-
- Moore T. S.; Winmill T. F. CLXXVII.—The State of Amines in Aqueous Solution. J. Chem. Soc., Trans. 1912, 101, 1635–1676. 10.1039/CT9120101635. - DOI
-
- Jorgensen W. L.; Pranata J. Importance of Secondary Interactions in Triply Hydrogen Bonded Complexes: Guanine-Cytosine vs Uracil-2,6-Diaminopyridine. J. Am. Chem. Soc. 1990, 112, 2008–2010. 10.1021/ja00161a061. - DOI
-
- Murray T. J.; Zimmerman S. C. New Triply Hydrogen Bonded Complexes with Highly Variable Stabilities. J. Am. Chem. Soc. 1992, 114, 4010–4011. 10.1021/ja00036a079. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

