PARP1: Structural insights and pharmacological targets for inhibition
- PMID: 33940558
- PMCID: PMC8206044
- DOI: 10.1016/j.dnarep.2021.103125
PARP1: Structural insights and pharmacological targets for inhibition
Abstract
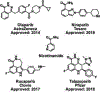
Poly(ADP-ribose) polymerase 1 (PARP1, also known as ADPRT1) is a multifunctional human ADP-ribosyltransferase. It plays a role in multiple DNA repair pathways, including the base excision repair (BER), non-homologous end joining (NHEJ), homologous recombination (HR), and Okazaki-fragment processing pathways. In response to DNA strand breaks, PARP1 covalently attaches ADP-ribose moieties to arginine, glutamate, aspartate, cysteine, lysine, and serine acceptor sites on both itself and other proteins. This signal recruits DNA repair proteins to the site of DNA damage. PARP1 binding to these sites enhances ADP-ribosylation via allosteric communication between the distant DNA binding and catalytic domains. In this review, we provide a general overview of PARP1 and emphasize novel potential approaches for pharmacological inhibition. Clinical PARP1 inhibitors bind the catalytic pocket, where they directly interfere with ADP-ribosylation. Some inhibitors may further enhance potency by "trapping" PARP1 on DNA via an allosteric mechanism, though this proposed mode of action remains controversial. PARP1 inhibitors are used clinically to treat some cancers, but resistance is common, so novel pharmacological approaches are urgently needed. One approach may be to design novel small molecules that bind at inter-domain interfaces that are essential for PARP1 allostery. To illustrate these points, this review also includes instructive videos showing PARP1 structures and mechanisms.
Keywords: Cancer; Chemotherapy; DNA repair; Drug resistance; PARP1; Review.
Copyright © 2021. Published by Elsevier B.V.
Conflict of interest statement
Declaration of Competing Interest
The authors report no declarations of interest.
Figures




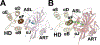

Similar articles
-
Serine-linked PARP1 auto-modification controls PARP inhibitor response.Nat Commun. 2021 Jul 1;12(1):4055. doi: 10.1038/s41467-021-24361-9. Nat Commun. 2021. PMID: 34210965 Free PMC article.
-
Histone ADP-ribosylation promotes resistance to PARP inhibitors by facilitating PARP1 release from DNA lesions.Proc Natl Acad Sci U S A. 2024 Jun 18;121(25):e2322689121. doi: 10.1073/pnas.2322689121. Epub 2024 Jun 12. Proc Natl Acad Sci U S A. 2024. PMID: 38865276 Free PMC article.
-
Medicinal chemistry approaches of poly ADP-Ribose polymerase 1 (PARP1) inhibitors as anticancer agents - A recent update.Eur J Med Chem. 2019 Mar 1;165:198-215. doi: 10.1016/j.ejmech.2019.01.024. Epub 2019 Jan 12. Eur J Med Chem. 2019. PMID: 30684797 Review.
-
Poly(ADP-ribose) Polymerase (PARP) and PARP Inhibitors: Mechanisms of Action and Role in Cardiovascular Disorders.Cardiovasc Toxicol. 2018 Dec;18(6):493-506. doi: 10.1007/s12012-018-9462-2. Cardiovasc Toxicol. 2018. PMID: 29968072 Review.
-
Captured snapshots of PARP1 in the active state reveal the mechanics of PARP1 allostery.Mol Cell. 2022 Aug 18;82(16):2939-2951.e5. doi: 10.1016/j.molcel.2022.06.011. Epub 2022 Jul 5. Mol Cell. 2022. PMID: 35793673 Free PMC article.
Cited by
-
Human papillomavirus E7 protein induces homologous recombination defects and PARPi sensitivity.J Cancer Res Clin Oncol. 2024 Jan 23;150(1):27. doi: 10.1007/s00432-023-05511-6. J Cancer Res Clin Oncol. 2024. PMID: 38263342 Free PMC article.
-
Expression of NORAD correlates with breast cancer aggressiveness and protects breast cancer cells from chemotherapy.Mol Ther Nucleic Acids. 2023 Aug 18;33:910-924. doi: 10.1016/j.omtn.2023.08.019. eCollection 2023 Sep 12. Mol Ther Nucleic Acids. 2023. PMID: 37680988 Free PMC article.
-
Poly(ADP-Ribose) Polymerase-1 Lacking Enzymatic Activity Is Not Compatible with Mouse Development.Cells. 2023 Aug 16;12(16):2078. doi: 10.3390/cells12162078. Cells. 2023. PMID: 37626888 Free PMC article.
-
Aminomethylmorpholino Nucleosides as Novel Inhibitors of PARP1 and PARP2: Experimental and Molecular Modeling Analyses of Their Selectivity and Mechanism of Action.Int J Mol Sci. 2024 Nov 22;25(23):12526. doi: 10.3390/ijms252312526. Int J Mol Sci. 2024. PMID: 39684238 Free PMC article.
-
Macrocarpal I induces immunogenic cell death and synergizes with immune checkpoint inhibition by targeting tubulin and PARP1 in colorectal cancer.Cell Death Discov. 2025 Feb 22;11(1):73. doi: 10.1038/s41420-025-02360-9. Cell Death Discov. 2025. PMID: 39987121 Free PMC article.
References
-
- Lin K-Y, Molecular Mechanism of Poly(ADP-ribosyl)ation Catalyzed by Human Poly(ADP-ribose) Polymerase-1.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

