A positive feedback loop reinforces the allergic immune response in human peanut allergy
- PMID: 33944900
- PMCID: PMC8103542
- DOI: 10.1084/jem.20201793
A positive feedback loop reinforces the allergic immune response in human peanut allergy
Abstract
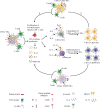
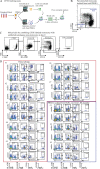
Food allergies are a leading cause of anaphylaxis, and cellular mechanisms involving antigen presentation likely play key roles in their pathogenesis. However, little is known about the response of specific antigen-presenting cell (APC) subsets to food allergens in the setting of food allergies. Here, we show that in peanut-allergic humans, peanut allergen drives the differentiation of CD209+ monocyte-derived dendritic cells (DCs) and CD23+ (FcєRII) myeloid dendritic cells through the action of allergen-specific CD4+ T cells. CD209+ DCs act reciprocally on the same peanut-specific CD4+ T cell population to reinforce Th2 cytokine expression in a positive feedback loop, which may explain the persistence of established food allergy. In support of this novel model, we show clinically that the initiation of oral immunotherapy (OIT) in peanut-allergic patients is associated with a decrease in CD209+ DCs, suggesting that breaking the cycle of positive feedback is associated with therapeutic effect.
© 2021 Zhou et al.
Conflict of interest statement
Disclosures: E.D. Mellins reported grants from Glaxo-Smith-Kline and Novartis outside the submitted work. K.C. Nadeau reported grants from the National Institute of Allergy and Infectious Diseases, the National Heart, Lung, and Blood Institute, the National Institute of Environmental Health Sciences, and Food Allergy Research and Education; "other" from World Allergy Organization, Cour Pharma, Before Brands, Alladapt, Latitude, IgGenix, Immune Tolerance Network, and the National Institutes of Health clinical research centers outside the submitted work. In addition, K.C. Nadeau had a patent to "inhibition of allergic reaction to peanut allergen using an IL-33 inhibitor" (US patent no. 62/647,389, filed 3/23/2018), a patent to "special oral formula for decreasing food allergy risk and treatment for food allergy (US patent no. 62/119,014, filed 2/20/15, issued 8/15/17) issued, a patent to "basophil activation-based diagnostic allergy test" (US application no. S10-392, filed 10/1/2010), a patent to "granulocyte-based methods for detecting and monitoring immune system disorders" (US application no. 12/686,121, filed 1/12/2010), a patent to "methods and assays for detecting and quantifying pure subpopulations of white blood cells in immune system disorders" (US patent no. 12/610,940, filed 11/2/2009, issued 8/12/2018), a patent to "mixed allergen compositions and methods for using the same" (US patent no. 10/064,936, issued 9/4/2018), and a patent to "microfluidic device and diagnostic methods for allergy testing based on detection of basophil activation" (US patent no. 62/767,444, filed 11/14/2018). No other disclosures were reported.
Figures












References
-
- Alonso, M.N., Wong M.T., Zhang A.L., Winer D., Suhoski M.M., Tolentino L.L., Gaitan J., Davidson M.G., Kung T.H., Galel D.M., et al. . 2011. T(H)1, T(H)2, and T(H)17 cells instruct monocytes to differentiate into specialized dendritic cell subsets. Blood. 118:3311–3320. 10.1182/blood-2011-03-341065 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

