DSCAM/PAK1 pathway suppression reverses neurogenesis deficits in iPSC-derived cerebral organoids from patients with Down syndrome
- PMID: 33945512
- PMCID: PMC8203468
- DOI: 10.1172/JCI135763
DSCAM/PAK1 pathway suppression reverses neurogenesis deficits in iPSC-derived cerebral organoids from patients with Down syndrome
Abstract
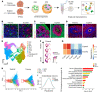
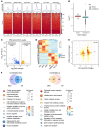
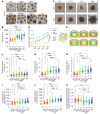
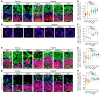
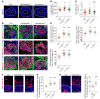
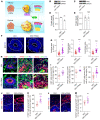
Down syndrome (DS), caused by trisomy of chromosome 21, occurs in 1 of every 800 live births. Early defects in cortical development likely account for the cognitive impairments in DS, although the underlying molecular mechanism remains elusive. Here, we performed histological assays and unbiased single-cell RNA-Seq (scRNA-Seq) analysis on cerebral organoids derived from 4 euploid cell lines and from induced pluripotent stem cells (iPSCs) from 3 individuals with trisomy 21 to explore cell-type-specific abnormalities associated with DS during early brain development. We found that neurogenesis was significantly affected, given the diminished proliferation and decreased expression of layer II and IV markers in cortical neurons in the subcortical regions; this may have been responsible for the reduced size of the organoids. Furthermore, suppression of the DSCAM/PAK1 pathway, which showed enhanced activity in DS, using CRISPR/Cas9, CRISPR interference (CRISPRi), or small-molecule inhibitor treatment reversed abnormal neurogenesis, thereby increasing the size of organoids derived from DS iPSCs. Our study demonstrates that 3D cortical organoids developed in vitro are a valuable model of DS and provide a direct link between dysregulation of the DSCAM/PAK1 pathway and developmental brain defects in DS.
Keywords: Embryonic stem cells; Neurodevelopment; Neuroscience; Stem cells; iPS cells.
Conflict of interest statement
Figures


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

