Copy Number Variation and Rearrangements Assessment in Cancer: Comparison of Droplet Digital PCR with the Current Approaches
- PMID: 33946969
- PMCID: PMC8124143
- DOI: 10.3390/ijms22094732
Copy Number Variation and Rearrangements Assessment in Cancer: Comparison of Droplet Digital PCR with the Current Approaches
Abstract
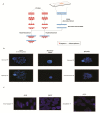
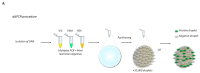
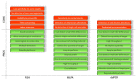
The cytogenetic and molecular assessment of deletions, amplifications and rearrangements are key aspects in the diagnosis and therapy of cancer. Not only the initial evaluation and classification of the disease, but also the follow-up of the tumor rely on these laboratory approaches. The therapeutic choice can be guided by the results of the laboratory testing. Genetic deletions and/or amplifications directly affect the susceptibility or the resistance to specific therapies. In an era of personalized medicine, the correct and reliable molecular characterization of the disease, also during the therapeutic path, acquires a pivotal role. Molecular assays like multiplex ligation-dependent probe amplification and droplet digital PCR represent exceptional tools for a sensitive and reliable detection of genetic alterations and deserve a role in molecular oncology. In this manuscript we provide a technical comparison of these two approaches with the golden standard represented by fluorescence in situ hybridization. We also describe some relevant targets currently evaluated with these techniques in solid and hematologic tumors.
Keywords: FISH; MLPA; copy number variation assessment; droplet digital PCR.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Figures


References
-
- Sussman R.T., Oran A.R., Paolillo C., Lieberman D., Morrissette J.J.D., Rosenbaum J.N. Validation of a next-generation sequencing assay targeting rna for the multiplexed detection of fusion transcripts and oncogenic isoforms. Arch. Pathol. Lab. Med. 2020;144:90–98. doi: 10.5858/arpa.2018-0441-OA. - DOI - PubMed
-
- Liu Y., Wu S., Shi X., Luo Y., Pang J., Wang C., Mao F., Liang Z., Zeng X. Her2 double-equivocal breast cancer in chinese patients: A high concordance of her2 status between different blocks from the same tumor. Breast Cancer Res. Treat. 2019;178:275–281. doi: 10.1007/s10549-019-05387-6. - DOI - PMC - PubMed
-
- Nevisi F., Yaghmaie M., Pashaiefar H., Alimoghaddam K., Iravani M., Javadi G., Ghavamzadeh A. Correlation of her2, mdm2, c-myc, c-met, and tp53 copy number alterations in circulating tumor cells with tissue in gastric cancer patients: A pilot study. Iran Biomed. J. 2020;24:47–53. doi: 10.29252/ibj.24.1.47. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

