Phylogroup stability contrasts with high within sequence type complex dynamics of Escherichia coli bloodstream infection isolates over a 12-year period
- PMID: 33952335
- PMCID: PMC8097792
- DOI: 10.1186/s13073-021-00892-0
Phylogroup stability contrasts with high within sequence type complex dynamics of Escherichia coli bloodstream infection isolates over a 12-year period
Abstract
Background: Escherichia coli is the leading cause of bloodstream infections, associated with a significant mortality. Recent genomic analyses revealed that few clonal lineages are involved in bloodstream infections and captured the emergence of some of them. However, data on within sequence type (ST) population genetic structure evolution are rare.
Methods: We compared whole genome sequences of 912 E. coli isolates responsible for bloodstream infections from two multicenter clinical trials that were conducted in the Paris area, France, 12 years apart, in teaching hospitals belonging to the same institution ("Assistance Publique-Hôpitaux de Paris"). We analyzed the strains at different levels of granularity, i.e., the phylogroup, the ST complex (STc), and the within STc clone taking into consideration the evolutionary history, the resistance, and virulence gene content as well as the antigenic diversity of the strains.
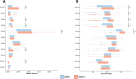
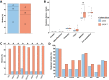
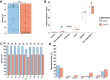
Results: We found a mix of stability and changes overtime, depending on the level of comparison. Overall, we observed an increase in antibiotic resistance associated to a restricted number of genetic determinants and in strain plasmidic content, whereas phylogroup distribution and virulence gene content remained constant. Focusing on STcs highlighted the pauci-clonality of the populations, with only 11 STcs responsible for more than 73% of the cases, dominated by five STcs (STc73, STc131, STc95, STc69, STc10). However, some STcs underwent dramatic variations, such as the global pandemic STc131, which replaced the previously predominant STc95. Moreover, within STc131, 95 and 69 genomic diversity analysis revealed a highly dynamic pattern, with reshuffling of the population linked to clonal replacement sometimes coupled with independent acquisitions of virulence factors such as the pap gene cluster bearing a papGII allele located on various pathogenicity islands. Additionally, STc10 exhibited huge antigenic diversity evidenced by numerous O:H serotype/fimH allele combinations, whichever the year of isolation.
Conclusions: Altogether, these data suggest that the bloodstream niche is occupied by a wide but specific phylogenetic diversity and that highly specialized extra-intestinal clones undergo frequent turnover at the within ST level. Additional worldwide epidemiological studies overtime are needed in different geographical and ecological contexts to assess how generalizable these data are.
Trial registration: ClinicalTrials.gov NCT02890901.
Keywords: Antibiotic resistance; Bloodstream infection; Clonal replacement; Escherichia coli; Extra-intestinal infection; Pandemic clones.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures




References
-
- Vihta K-D, Stoesser N, Llewelyn MJ, Quan TP, Davies T, Fawcett NJ, Dunn L, Jeffery K, Butler CC, Hayward G, Andersson M, Morgan M, Oakley S, Mason A, Hopkins S, Wyllie DH, Crook DW, Wilcox MH, Johnson AP, Peto TEA, Walker AS. Trends over time in Escherichia coli bloodstream infections, urinary tract infections, and antibiotic susceptibilities in Oxfordshire, UK, 1998–2016: a study of electronic health records. Lancet Infect Dis. 2018;18(10):1138–1149. doi: 10.1016/S1473-3099(18)30353-0. - DOI - PMC - PubMed
-
- Lefort A, Panhard X, Clermont O, Woerther P-L, Branger C, Mentre F, Fantin B, Wolff M, Denamur E, for the COLIBAFI Group Host factors and portal of entry outweigh bacterial determinants to predict the severity of Escherichia coli bacteremia. Journal of Clinical Microbiology. 2011;49(3):777–783. doi: 10.1128/JCM.01902-10. - DOI - PMC - PubMed
-
- Yoon E-J, Choi MH, Park YS, Lee HS, Kim D, Lee H, Shin KS, Shin JH, Uh Y, Kim YA, Shin JH, Jeong SH. Impact of host-pathogen-treatment tripartite components on early mortality of patients with Escherichia coli bloodstream infection: prospective observational study. EBioMedicine. 2018;35:76–86. doi: 10.1016/j.ebiom.2018.08.029. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Associated data
LinkOut - more resources
Full Text Sources
Medical
Research Materials

