Identification of seven tumor-educated platelets RNAs for cancer diagnosis
- PMID: 33955587
- PMCID: PMC8183939
- DOI: 10.1002/jcla.23791
Identification of seven tumor-educated platelets RNAs for cancer diagnosis
Abstract
Background: Tumor-educated platelets (TEPs) may enable blood-based cancer diagnosis. This study aimed to identify diagnostic TEPs genes involved in carcinogenesis.
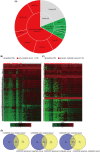
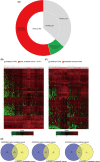
Materials and methods: The TEPs differentially expressed genes (DEGs) between healthy samples and early/advanced cancer samples were obtained using bioinformatics. Gene ontology (GO) analysis and Kyoto encyclopedia of genes and genomes (KEGG) pathway enrichment analysis were used to identify the pathways and functional annotation of TEPs DEGs. Protein-protein interaction of these TEPs DEGs was analyzed based on the STRING database and visualized by Cytoscape software. The correlation analysis and diagnostic analysis were performed to evaluate the diagnostic value of TEPs mRNAs expression for early/advanced cancers. Quantitative real-time PCR (qRT-PCR) was applied to validate the role of DEGs in cancers.
Results: TEPs mRNAs were mostly involved in protein binding, extracellular matrix, and cellular protein metabolic process. RSL24D1 was negatively correlated to early-stage cancers compared to healthy controls and may be potentially used for early cancer diagnosis. In addition, HPSE, IFI27, LGALS3BP, CRYM, HBD, COL6A3, LAMB2, and IFITM3 showed an upward trend in the expression from early to advanced cancer stages. Moreover, ARL2, FCGR2A, and KLHDC8B were positively associated with advanced, metastatic cancers compared to healthy controls. Among the 12 selected DEGs, the expression of 7 DEGs, including RSL24D1, IFI27, CRYM, HBD, IFITM3, FCGR2A, and KLHDC8B, were verified by the qRT-PCR method.
Conclusion: This study suggests that the 7-gene TEPs liquid-biopsy biomarkers may be used for cancer diagnosis and monitoring.
Keywords: bioinformatics analysis; diagnosis; mRNA; tumor educated platelets.
© 2021 The Authors. Journal of Clinical Laboratory Analysis published by Wiley Periodicals LLC.
Conflict of interest statement
The authors declare no conflict of interest.
Figures





References
-
- Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020;70(1):7‐30. - PubMed
-
- Vaidyanathan R, Soon RH, Zhang P, Jiang K, Lim CT. Cancer diagnosis: from tumor to liquid biopsy and beyond. Lab Chip. 2018;19(1):11‐34. - PubMed
-
- Krebs MG, Metcalf RL, Carter L, Brady G, Blackhall FH, Dive C. Molecular analysis of circulating tumour cells‐biology and biomarkers. Nat Rev Clin Oncol. 2014;11(3):129‐144. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

