Establishment of an Agrobacterium-mediated genetic transformation and CRISPR/Cas9-mediated targeted mutagenesis in Hemp (Cannabis Sativa L.)
- PMID: 33960612
- PMCID: PMC8486249
- DOI: 10.1111/pbi.13611
Establishment of an Agrobacterium-mediated genetic transformation and CRISPR/Cas9-mediated targeted mutagenesis in Hemp (Cannabis Sativa L.)
Abstract
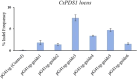
Hemp (Cannabis sativa L.) is an annual and typically dioecious crop. Due to the therapeutic potential for human diseases, phytocannabinoids as a medical therapy is getting more attention recently. Several candidate genes involved in cannabinoid biosynthesis have been elucidated using omics analysis. However, the gene function was not fully validated due to few reports of stable transformation for Cannabis tissues. In this study, we firstly report the successful generation of gene-edited plants using an Agrobacterium-mediated transformation method in C. sativa. DMG278 achieved the highest shoot induction rate, which was selected as the model strain for transformation. By overexpressing the cannabis developmental regulator chimera in the embryo hypocotyls of immature grains, the shoot regeneration efficiency was substantially increased. We used CRISPR/Cas9 technology to edit the phytoene desaturase gene and finally generated four edited cannabis seedlings with albino phenotype. Moreover, we propagated the transgenic plants and validated the stable integration of T-DNA in cannabis genome.
Keywords: Agrobacterium-mediated transformation; CRISPR; Cas9; Hemp; genome editing.
© 2021 The Authors. Plant Biotechnology Journal published by Society for Experimental Biology and The Association of Applied Biologists and John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no conflict of interest.
Figures





Similar articles
-
Optimizing Agrobacterium-Mediated Transformation and CRISPR-Cas9 Gene Editing in the tropical japonica Rice Variety Presidio.Int J Mol Sci. 2021 Oct 9;22(20):10909. doi: 10.3390/ijms222010909. Int J Mol Sci. 2021. PMID: 34681568 Free PMC article.
-
An efficient and specific CRISPR-Cas9 genome editing system targeting soybean phytoene desaturase genes.BMC Biotechnol. 2022 Feb 15;22(1):7. doi: 10.1186/s12896-022-00737-7. BMC Biotechnol. 2022. PMID: 35168613 Free PMC article.
-
Establishment of CRISPR/Cas9 mediated targeted mutagenesis in hop (Humulus lupulus).Plant Physiol Biochem. 2021 Mar;160:1-7. doi: 10.1016/j.plaphy.2021.01.006. Epub 2021 Jan 7. Plant Physiol Biochem. 2021. PMID: 33445042
-
Advances and Perspectives in Tissue Culture and Genetic Engineering of Cannabis.Int J Mol Sci. 2021 May 26;22(11):5671. doi: 10.3390/ijms22115671. Int J Mol Sci. 2021. PMID: 34073522 Free PMC article. Review.
-
Agrobacterium-Mediated Transformation for the Development of Transgenic Crops; Present and Future Prospects.Mol Biotechnol. 2024 Aug;66(8):1836-1852. doi: 10.1007/s12033-023-00826-8. Epub 2023 Aug 13. Mol Biotechnol. 2024. PMID: 37573566 Review.
Cited by
-
Multi-Omics Approaches to Study Molecular Mechanisms in Cannabis sativa.Plants (Basel). 2022 Aug 22;11(16):2182. doi: 10.3390/plants11162182. Plants (Basel). 2022. PMID: 36015485 Free PMC article. Review.
-
New Insight into Ornamental Applications of Cannabis: Perspectives and Challenges.Plants (Basel). 2022 Sep 13;11(18):2383. doi: 10.3390/plants11182383. Plants (Basel). 2022. PMID: 36145783 Free PMC article.
-
In planta Female Flower Agroinfiltration Alters the Cannabinoid Composition in Industrial Hemp (Cannabis sativa L.).Front Plant Sci. 2022 Jul 21;13:921970. doi: 10.3389/fpls.2022.921970. eCollection 2022. Front Plant Sci. 2022. PMID: 35941940 Free PMC article.
-
Improving Sedum plumbizincicola genetic transformation with the SpGRF4-SpGIF1 gene and the self-excision CRE/LoxP system.Planta. 2024 Apr 9;259(5):119. doi: 10.1007/s00425-024-04393-3. Planta. 2024. PMID: 38594473
-
Challenges and potentials of new breeding techniques in Cannabis sativa.Front Plant Sci. 2023 Jun 8;14:1154332. doi: 10.3389/fpls.2023.1154332. eCollection 2023. Front Plant Sci. 2023. PMID: 37360738 Free PMC article. Review.
References
-
- Ahmed, S. , Gao, X. , Jahan, M.A. , Adams, M. , Wu, N. and Kovinich, N. (2021) Nanoparticle‐based genetic transformation of Cannabis sativa . J. Biotechnol. 326, 48–51. - PubMed
-
- Borgato, L. , Pisani, F. and Furini, A. (2007) Plant regeneration from leaf protoplasts of Solanum virginianum L. (Solanaceae). Plant Cell Tissue Organ Cult. 88, 247–252.
-
- Cheng, C. , Zang, G. , Zhao, L. , Gao, C. , Tang, Q. , Chen, J. , Guo, X. et al. (2016) A rapid shoot regeneration protocol from the cotyledons of hemp (Cannabis sativa L.). Ind. Crops Prod. 83, 61–65.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

