Construction of Mycoplasma hyopneumoniae P97 Null Mutants
- PMID: 33967967
- PMCID: PMC8101707
- DOI: 10.3389/fmicb.2021.518791
Construction of Mycoplasma hyopneumoniae P97 Null Mutants
Abstract
Mycoplasma hyopneumoniae is the causative agent of enzootic pneumonia, a world-wide problem in the pig industry. This disease is characterized by a dry, non-productive cough, labored breathing, and pneumonia. Despite years of research, vaccines are marginally effective, and none fully protect pigs in a production environment. A better understanding of the host-pathogen interactions of the M. hyopneumoniae-pig disease, which are complex and involve both host and pathogen components, is required. Among the surface proteins involved in virulence are members of two gene families called P97 and P102. These proteins are the adhesins directing attachment of the organism to the swine respiratory epithelium. P97 is the major ciliary binding adhesin and has been studied extensively. Monoclonal antibodies that block its binding to swine cilia have contributed extensively to its characterization. In this study we use recombination to construct null mutants of P97 in M. hyopneumoniae and characterize the resulting mutants in terms of loss of protein by immunoblot using monoclonal antibodies, ability to bind purified swine cilia, and adherence to PK15 cells. Various approaches to recombination with this fastidious mycoplasma were tested including intact plasmid DNA, single-stranded DNA, and linear DNA with and without a heterologous RecA protein. Our results indicate that recombination can be used to generate site-specific mutants in M. hyopneumoniae. P97 mutants are deficient in cilia binding and PK15 cell adherence, and lack the characteristic banding pattern seen in immunoblots developed with the anti-P97 monoclonal antibody.
Keywords: Mycoplasma hyopneumoniae; P97; adherence assay; adherence protein; immunoblot; null mutant; recombination.
Copyright © 2021 Clampitt, Madsen and Minion.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


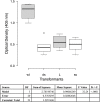


References
-
- Bogema D. R., Deutscher A. T., Woolley L. K., Seymour L. M., Raymond B. B., Tacchi J. L., et al. (2012). Characterization of cleavage events in the multifunctional cilium adhesin Mhp684 (P146) reveals a mechanism by which Mycoplasma hyopneumoniae regulates surface topography. mBio 3:e00282-11. - PMC - PubMed
-
- Bogema D. R., Scott N. E., Padula M., Tacchi J. L., Jenkins C., Cordwell S. J., et al. (2011). Sequence TTKF | QE defines the site of proteolytic cleavage in Mhp683 protein, a novel glycosaminoglycan and cilium adhesin of Mycoplasma hyopneumoniae. J. Biol. Chem. 286 41217–41229. 10.1074/jbc.m111.226084 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources

