Endogenous Retroviral-K Envelope Is a Novel Tumor Antigen and Prognostic Indicator of Renal Cell Carcinoma
- PMID: 33968761
- PMCID: PMC8100683
- DOI: 10.3389/fonc.2021.657187
Endogenous Retroviral-K Envelope Is a Novel Tumor Antigen and Prognostic Indicator of Renal Cell Carcinoma
Abstract
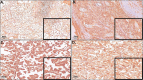
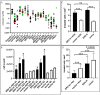
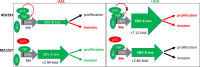
Renal cell carcinoma (RCC) is one of the ten most common cancers for men and women with an approximate 75% overall 5-year survival. Sixteen histological tumor subtypes exist and the most common are papillary, chromophobe and clear cell renal cell carcinoma (ccRCC) representing 85% of all RCC. Although epigenetically silenced, endogenous retroviral (ERV) genes become activated in tumors and function to ignite immune responses. Research has intensified to understand ERV protein function and their role as tumor antigens and targets for cancer (immune) therapy. ERV-K env is overexpressed and implicated as a therapeutic target for breast cancer, however studies in RCC are limited. In this investigation a human RCC tissue microarray (TMA) (n=374) predominantly consisting of the most common histological tumor subtypes was hybridized with an ERV-K env antibody and correlated with patient clinical data. TMA results showed the highest amount of ERV-K env protein expression and the strongest significant membrane expression in ccRCC versus other RCC subtypes. High ERV-K env total protein expression of all tumor subtypes significantly correlated with low tumor grading and a longer disease specific survival using multivariable analyses. Cell proliferation and invasion were assayed using the kidney cell lines HEK293 with wild-type p53 and a ccRCC cell line MZ1257RC mutated for p53. Transfecting these cell lines with a codon optimized ERV-K113 env overexpressing CMV vector was performed with or without 5'-Aza-2'-deoxycytidine (Aza) treatment to sustain promoter de-methylation. MZ1257RC showed induction of ERV-K113 expression and significantly increased both proliferation and invasion in the presence or absence of Aza. HEK293 cells demonstrated a restriction of ERV-K113 env expression and invasion with no changes in proliferation in the absence of Aza. However, in the presence of Aza despite increased ERV-K113 env expression, an inhibition of HEK293 proliferation and a further restriction of invasion was found. This study supports ERV-K env as a single prognostic indicator for better survival of RCC, which we propose represents a new tumor antigen. In addition, ERV-K env significantly regulates proliferation and invasion depending on p53 status and Aza treatment.
Keywords: ERV-K; azacytidine; endogenous retrovirus; invasion; p53; patient prognosis; renal cell carcinoma; tumor antigen.
Copyright © 2021 Weyerer, Strissel, Stöhr, Eckstein, Wach, Taubert, Brandl, Geppert, Wullich, Cynis, Beckmann, Seliger, Hartmann and Strick.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



References
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

