X-Ray Photoelectron Spectroscopy on Microbial Cell Surfaces: A Forgotten Method for the Characterization of Microorganisms Encapsulated With Surface-Engineered Shells
- PMID: 33968904
- PMCID: PMC8100684
- DOI: 10.3389/fchem.2021.666159
X-Ray Photoelectron Spectroscopy on Microbial Cell Surfaces: A Forgotten Method for the Characterization of Microorganisms Encapsulated With Surface-Engineered Shells
Abstract
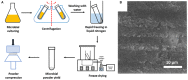
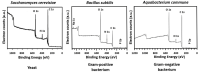
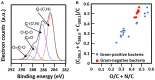
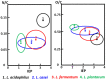
Encapsulation of single microbial cells by surface-engineered shells has great potential for the protection of yeasts and bacteria against harsh environmental conditions, such as elevated temperatures, UV light, extreme pH values, and antimicrobials. Encapsulation with functionalized shells can also alter the surface characteristics of cells in a way that can make them more suitable to perform their function in complex environments, including bio-reactors, bio-fuel production, biosensors, and the human body. Surface-engineered shells bear as an advantage above genetically-engineered microorganisms that the protection and functionalization added are temporary and disappear upon microbial growth, ultimately breaking a shell. Therewith, the danger of creating a "super-bug," resistant to all known antimicrobial measures does not exist for surface-engineered shells. Encapsulating shells around single microorganisms are predominantly characterized by electron microscopy, energy-dispersive X-ray spectroscopy, Fourier transform infrared spectroscopy, particulate micro-electrophoresis, nitrogen adsorption-desorption isotherms, and X-ray diffraction. It is amazing that X-ray Photoelectron Spectroscopy (XPS) is forgotten as a method to characterize encapsulated yeasts and bacteria. XPS was introduced several decades ago to characterize the elemental composition of microbial cell surfaces. Microbial sample preparation requires freeze-drying which leaves microorganisms intact. Freeze-dried microorganisms form a powder that can be easily pressed in small cups, suitable for insertion in the high vacuum of an XPS machine and obtaining high resolution spectra. Typically, XPS measures carbon, nitrogen, oxygen and phosphorus as the most common elements in microbial cell surfaces. Models exist to transform these compositions into well-known, biochemical cell surface components, including proteins, polysaccharides, chitin, glucan, teichoic acid, peptidoglycan, and hydrocarbon like components. Moreover, elemental surface compositions of many different microbial strains and species in freeze-dried conditions, related with zeta potentials of microbial cells, measured in a hydrated state. Relationships between elemental surface compositions measured using XPS in vacuum with characteristics measured in a hydrated state have been taken as a validation of microbial cell surface XPS. Despite the merits of microbial cell surface XPS, XPS has seldom been applied to characterize the many different types of surface-engineered shells around yeasts and bacteria currently described in the literature. In this review, we aim to advocate the use of XPS as a forgotten method for microbial cell surface characterization, for use on surface-engineered shells encapsulating microorganisms.
Keywords: beer brewing; biomineralization; biosorption; click chemistry; nanobiomaterials; self-assembly; yolk shell; zeta potentials.
Copyright © 2021 Wei, Yang, van der Mei and Busscher.
Conflict of interest statement
HB is also director of a consulting company, SASA BV (GN Schutterlaan 4, 9797 PC Thesinge, Netherlands). The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures




Similar articles
-
Planning Implications Related to Sterilization-Sensitive Science Investigations Associated with Mars Sample Return (MSR).Astrobiology. 2022 Jun;22(S1):S112-S164. doi: 10.1089/AST.2021.0113. Epub 2022 May 19. Astrobiology. 2022. PMID: 34904892
-
Interfacial interactions between protective, surface-engineered shells and encapsulated bacteria with different cell surface composition.Nanoscale. 2021 Apr 21;13(15):7220-7233. doi: 10.1039/d0nr09204e. Epub 2021 Apr 7. Nanoscale. 2021. PMID: 33889889
-
The use of X-ray photoelectron spectroscopy for the study of oral streptococcal cell surfaces.Adv Dent Res. 1997 Nov;11(4):388-94. doi: 10.1177/08959374970110040301. Adv Dent Res. 1997. PMID: 9470495
-
Low-voltage SEM of air-sensitive powders: From sample preparation to micro/nano analysis with secondary electron hyperspectral imaging.Micron. 2022 May;156:103234. doi: 10.1016/j.micron.2022.103234. Epub 2022 Mar 14. Micron. 2022. PMID: 35325668 Review.
-
Upflow anaerobic sludge blanket reactor--a review.Indian J Environ Health. 2001 Apr;43(2):1-82. Indian J Environ Health. 2001. PMID: 12397675 Review.
Cited by
-
Activation of a passive, mesoporous silica nanoparticle layer through attachment of bacterially-derived carbon-quantum-dots for protection and functional enhancement of probiotics.Mater Today Bio. 2022 May 17;15:100293. doi: 10.1016/j.mtbio.2022.100293. eCollection 2022 Jun. Mater Today Bio. 2022. PMID: 35634173 Free PMC article.
-
Microbiologically Influenced Corrosion of Q235 Carbon Steel by Ectothiorhodospira sp.Int J Environ Res Public Health. 2022 Nov 21;19(22):15416. doi: 10.3390/ijerph192215416. Int J Environ Res Public Health. 2022. PMID: 36430135 Free PMC article.
-
Inheritance of physico-chemical properties and ROS generation by carbon quantum dots derived from pyrolytically carbonized bacterial sources.Mater Today Bio. 2021 Oct 15;12:100151. doi: 10.1016/j.mtbio.2021.100151. eCollection 2021 Sep. Mater Today Bio. 2021. PMID: 34746735 Free PMC article.
-
Advancing modified biochar for sustainable agriculture: a comprehensive review on characterization, analysis, and soil performance.Biochar. 2025;7(1):8. doi: 10.1007/s42773-024-00397-0. Epub 2025 Jan 3. Biochar. 2025. PMID: 39758611 Free PMC article. Review.
References
-
- Amory D. E., Rouxhet P. G. (1988a). Surface properties of Saccharomyces cerevisiae and Saccharomyces carlsbergensis: chemical composition, electrostatic charge, and hydrophobicity. Biochim. Biophys. Acta 938, 61–79. 10.1016/0005-2736(88)90122-8 - DOI
-
- Amory D. E., Rouxhet P. G. (1988b). Flocculence of brewery yeasts and their surface properties: chemical composition, electrostatic charge, and hydrophobicity. J. Inst. Brew. 94, 79–84. 10.1002/j.2050-0416.1988.tb04561.x - DOI
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources

