Sensitivity to expression levels underlies differential dominance of a putative null allele of the Drosophila tβh gene in behavioral phenotypes
- PMID: 33970909
- PMCID: PMC8136860
- DOI: 10.1371/journal.pbio.3001228
Sensitivity to expression levels underlies differential dominance of a putative null allele of the Drosophila tβh gene in behavioral phenotypes
Abstract
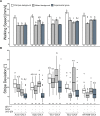
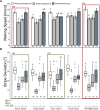
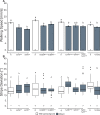
The biogenic amine octopamine (OA) and its precursor tyramine (TA) are involved in controlling a plethora of different physiological and behavioral processes. The tyramine-β-hydroxylase (tβh) gene encodes the enzyme catalyzing the last synthesis step from TA to OA. Here, we report differential dominance (from recessive to overdominant) of the putative null tβhnM18 allele in 2 behavioral measures in Buridan's paradigm (walking speed and stripe deviation) and in proboscis extension (sugar sensitivity) in the fruit fly Drosophila melanogaster. The behavioral analysis of transgenic tβh expression experiments in mutant and wild-type flies as well as of OA and TA receptor mutants revealed a complex interaction of both aminergic systems. Our analysis suggests that the different neuronal networks responsible for the 3 phenotypes show differential sensitivity to tβh gene expression levels. The evidence suggests that this sensitivity is brought about by a TA/OA opponent system modulating the involved neuronal circuits. This conclusion has important implications for standard transgenic techniques commonly used in functional genetics.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



References
-
- Fisher RA. The genetical theory of natural selection. Oxford: Oxford University Press; 1930.
-
- Waxman D, Peck JR. Pleiotropy and the preservation of perfection. Science. 1998;279:1210–3. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

