Associative Learning Requires Neurofibromin to Modulate GABAergic Inputs to Drosophila Mushroom Bodies
- PMID: 33972401
- PMCID: PMC8211548
- DOI: 10.1523/JNEUROSCI.1605-20.2021
Associative Learning Requires Neurofibromin to Modulate GABAergic Inputs to Drosophila Mushroom Bodies
Abstract
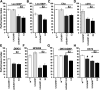
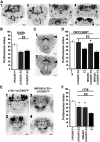
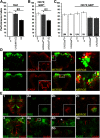
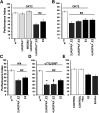
Cognitive dysfunction is among the hallmark symptoms of Neurofibromatosis 1, and accordingly, loss of the Drosophila melanogaster ortholog of Neurofibromin 1 (dNf1) precipitates associative learning deficits. However, the affected circuitry in the adult CNS remained unclear and the compromised mechanisms debatable. Although the main evolutionarily conserved function attributed to Nf1 is to inactivate Ras, decreased cAMP signaling on its loss has been thought to underlie impaired learning. Using mixed sex populations, we determine that dNf1 loss results in excess GABAergic signaling to the central for associative learning mushroom body (MB) neurons, apparently suppressing learning. dNf1 is necessary and sufficient for learning within these non-MB neurons, as a dAlk and Ras1-dependent, but PKA-independent modulator of GABAergic neurotransmission. Surprisingly, we also uncovered and discuss a postsynaptic Ras1-dependent, but dNf1-independnet signaling within the MBs that apparently responds to presynaptic GABA levels and contributes to the learning deficit of the mutants.
Keywords: Drosophila; GABA; Ras1; cAMP; learning; neurofibromatosis 1.
Copyright © 2021 the authors.
Conflict of interest statement
The authors declare no competing financial interests.
Figures

Similar articles
-
The receptor tyrosine kinase Alk controls neurofibromin functions in Drosophila growth and learning.PLoS Genet. 2011 Sep;7(9):e1002281. doi: 10.1371/journal.pgen.1002281. Epub 2011 Sep 15. PLoS Genet. 2011. PMID: 21949657 Free PMC article.
-
Neurofibromin 1 in mushroom body neurons mediates circadian wake drive through activating cAMP-PKA signaling.Nat Commun. 2021 Oct 1;12(1):5758. doi: 10.1038/s41467-021-26031-2. Nat Commun. 2021. PMID: 34599173 Free PMC article.
-
Genetic and functional studies implicate synaptic overgrowth and ring gland cAMP/PKA signaling defects in the Drosophila melanogaster neurofibromatosis-1 growth deficiency.PLoS Genet. 2013 Nov;9(11):e1003958. doi: 10.1371/journal.pgen.1003958. Epub 2013 Nov 21. PLoS Genet. 2013. PMID: 24278035 Free PMC article.
-
Suppression of inhibitory GABAergic transmission by cAMP signaling pathway: alterations in learning and memory mutants.Eur J Neurosci. 2013 May;37(9):1383-93. doi: 10.1111/ejn.12144. Epub 2013 Feb 7. Eur J Neurosci. 2013. PMID: 23387411 Free PMC article.
-
Bridging behavior and physiology: ion-channel perspective on mushroom body-dependent olfactory learning and memory in Drosophila.J Cell Physiol. 2006 Dec;209(3):1046-53. doi: 10.1002/jcp.20764. J Cell Physiol. 2006. PMID: 16924658 Review.
Cited by
-
Modeling neurodegenerative and neurodevelopmental disorders in the Drosophila mushroom body.Learn Mem. 2024 Jun 14;31(5):a053816. doi: 10.1101/lm.053816.123. Print 2024 May. Learn Mem. 2024. PMID: 38876485 Free PMC article. Review.
-
Loss of NF1 in Drosophila Larvae Causes Tactile Hypersensitivity and Impaired Synaptic Transmission at the Neuromuscular Junction.J Neurosci. 2022 Dec 14;42(50):9450-9472. doi: 10.1523/JNEUROSCI.0562-22.2022. Epub 2022 Nov 7. J Neurosci. 2022. PMID: 36344265 Free PMC article.
-
Neurofibromin 1 mediates sleep depth in Drosophila.PLoS Genet. 2023 Dec 13;19(12):e1011049. doi: 10.1371/journal.pgen.1011049. eCollection 2023 Dec. PLoS Genet. 2023. PMID: 38091360 Free PMC article.
-
Drosophila Contributions towards Understanding Neurofibromatosis 1.Cells. 2024 Apr 21;13(8):721. doi: 10.3390/cells13080721. Cells. 2024. PMID: 38667335 Free PMC article. Review.
-
Neurofibromin deficiency alters the patterning and prioritization of motor behaviors in a state-dependent manner.bioRxiv [Preprint]. 2024 Aug 9:2024.08.08.607070. doi: 10.1101/2024.08.08.607070. bioRxiv. 2024. Update in: J Neurosci. 2025 Apr 16;45(16):e1531242025. doi: 10.1523/JNEUROSCI.1531-24.2025. PMID: 39149363 Free PMC article. Updated. Preprint.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous
