The Transposable Element Environment of Human Genes Differs According to Their Duplication Status and Essentiality
- PMID: 33973013
- PMCID: PMC8155550
- DOI: 10.1093/gbe/evab062
The Transposable Element Environment of Human Genes Differs According to Their Duplication Status and Essentiality
Erratum in
-
Erratum to: The Transposable Element Environment of Human Genes Differs According to Their Duplication Status and Essentiality.Genome Biol Evol. 2021 Sep 1;13(9):evab175. doi: 10.1093/gbe/evab175. Genome Biol Evol. 2021. PMID: 34508264 Free PMC article. No abstract available.
Abstract
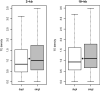
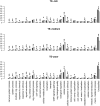
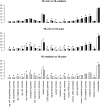
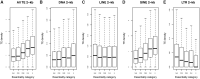
Transposable elements (TEs) are major components of eukaryotic genomes and represent approximately 45% of the human genome. TEs can be important sources of novelty in genomes and there is increasing evidence that TEs contribute to the evolution of gene regulation in mammals. Gene duplication is an evolutionary mechanism that also provides new genetic material and opportunities to acquire new functions. To investigate how duplicated genes are maintained in genomes, here, we explored the TE environment of duplicated and singleton genes. We found that singleton genes have more short-interspersed nuclear elements and DNA transposons in their vicinity than duplicated genes, whereas long-interspersed nuclear elements and long-terminal repeat retrotransposons have accumulated more near duplicated genes. We also discovered that this result is highly associated with the degree of essentiality of the genes with an unexpected accumulation of short-interspersed nuclear elements and DNA transposons around the more-essential genes. Our results underline the importance of taking into account the TE environment of genes to better understand how duplicated genes are maintained in genomes.
Keywords: LINE; SINE; essential genes; gene duplication; gene evolution; transposable elements.
© The Author(s) 2021. Published by Oxford University Press on behalf of the Society for Molecular Biology and Evolution.
Figures
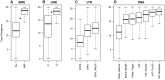
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

