Expression Profiles of Digestive Genes in the Gut and Salivary Glands of Tarnished Plant Bug (Hemiptera: Miridae)
- PMID: 33974083
- PMCID: PMC8112305
- DOI: 10.1093/jisesa/ieab028
Expression Profiles of Digestive Genes in the Gut and Salivary Glands of Tarnished Plant Bug (Hemiptera: Miridae)
Abstract
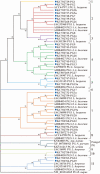
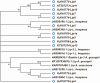
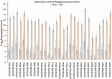
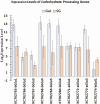
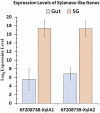
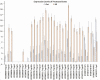
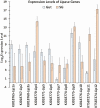
Host plant preference of agricultural pests may shift throughout the growing season, allowing the pests to persist on wild hosts when crops are not available. Lygus Hahn (Hemiptera: Miridae) bugs are severe pests of cotton during flowering and fruiting stages, but can persist on alternative crops, or on weed species. Diversity of digestive enzymes produced by salivary glands and gut tissues play a pivotal role in an organism's ability to utilize various food sources. Polyphagous insects produce an array of enzymes that can process carbohydrates, lipids, and proteins. In this study, the digestive enzyme repertoire of the tarnished plant bug, Lygus lineolaris (Palisot de Beauvois), was identified by high-throughput sequencing followed by cDNA cloning and sequencing. This study identified 87 digestive genes, including 30 polygalacturonases (PG), one β-galactosidase, three α-glucosidases, six β-glucosidases, 28 trypsin-like proteases, three serine proteases, one apyrase-like protease, one cysteine protease, 12 lipases, and two transcripts with low similarity to a xylanase A-like genes. RNA-Seq expression profiles of these digestive genes in adult tarnished plant bugs revealed that 57 and 12 genes were differentially expressed in the salivary gland and gut (≥5-fold, P ≤ 0.01), respectively. All polygalacturonase genes, most proteases, and two xylanase-like genes were differentially expressed in salivary glands, while most of the carbohydrate and lipid processing enzymes were differentially expressed in the gut. Seven of the proteases (KF208689, KF208697, KF208698, KF208699, KF208700, KF208701, and KF208702) were not detected in either the gut or salivary glands.
Keywords: RNA-Seq; digestive enzyme; expression profile; salivary gland; tarnished plant bug.
Published by Oxford University Press on behalf of Entomological Society of America 2021.
Figures


References
-
- Allen, M. L. 2007. Expressed sequenced tags from Lygus lineolaris (Hemiptera: Miridae), the tarnished plant bug. Genet. Mol. Res. 6: 206–213. - PubMed
-
- Allen, M. L., and W. B.Walker, 3rd. 2012. Saliva of Lygus lineolaris digests double stranded ribonucleic acids. J. Insect Physiol. 58: 391–396. - PubMed
-
- Almagro Armenteros, J. J., Tsirigos K. D., Sønderby C. K., Petersen T. N., Winther O., Brunak S., von Heijne G., and Nielsen H.. . 2019. SignalP 5.0 improves signal peptide predictions using deep neural networks. Nat. Biotechnol. 37: 420–423. - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

