Variation in selective constraints along the Plasmodium life cycle
- PMID: 33975022
- PMCID: PMC8205998
- DOI: 10.1016/j.meegid.2021.104908
Variation in selective constraints along the Plasmodium life cycle
Abstract
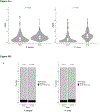
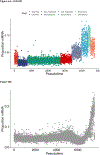
Plasmodium parasites, the cause of malaria, have a complex life cycle, infecting alternatively vertebrate hosts and female Anopheles mosquitoes and undergoing intra- and extra-cellular development in several organs of these hosts. Most of the ~5000 protein-coding genes present in Plasmodium genomes are only expressed at specific life stages, and different genes might therefore be subject to different selective pressures depending on the biological activity of the parasite and its microenvironment at this point in development. Here, we estimate the selective constraints on the protein-coding sequences of all annotated genes of rodent and primate Plasmodium parasites and, using data from scRNA-seq experiments spanning many developmental stages, analyze their variation with regard to when these genes are expressed in the parasite life cycle. Our study reveals extensive variation in selective constraints throughout the parasites' development and highlights stages that are evolving more rapidly than others. These findings provide novel insights into the biology of these parasites and could provide important information to develop better treatment strategies or vaccines against these medically-important organisms.
Keywords: Plasmodium; Selection; scRNA-seq.
Copyright © 2021 The Authors. Published by Elsevier B.V. All rights reserved.
Figures




References
-
- Amambua-Ngwa A, Tetteh KKA, Manske M, Gomez-Escobar N, Stewart LB, Deerhake ME, Cheeseman IH, Newbold CI, Holder AA, Knuepfer E, et al. 2012. Population Genomic Scan for Candidate Signatures of Balancing Selection to Guide Antigen Characterization in Malaria Parasites. PLoS Genet. 8(11). doi:10.1371/journal.pgen.1002992. - DOI - PMC - PubMed
-
- Auburn S, Böhme U, Steinbiss S, Trimarsanto H, Hostetler J, Sanders M, Gao Q, Nosten F, Newbold CI, Berriman M, et al. 2016. A new Plasmodium vivax reference sequence with improved assembly of the subtelomeres reveals an abundance of pir genes. Wellcome Open Res. 1(0):1–12. doi:10.12688/wellcomeopenres.9876.1. - DOI - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

