A Metabolite of Pseudomonas Triggers Prophage-Selective Lysogenic to Lytic Conversion in Staphylococcus aureus
- PMID: 33978401
- PMCID: PMC8193634
- DOI: 10.1021/jacs.1c01275
A Metabolite of Pseudomonas Triggers Prophage-Selective Lysogenic to Lytic Conversion in Staphylococcus aureus
Abstract
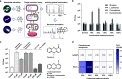
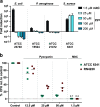
Bacteriophages have major impact on their microbial hosts and shape entire microbial communities. The majority of these phages are latent and reside as prophages integrated in the genomes of their microbial hosts. A variety of intricate regulatory systems determine the switch from a lysogenic to lytic life style, but so far strategies are lacking to selectively control prophage induction by small molecules. Here we show that Pseudomonas aeruginosa deploys a trigger factor to hijack the lysogenic to lytic switch of a polylysogenic Staphylococcus aureus strain causing the selective production of only one of its prophages. Fractionating extracts of P. aeruginosa identified the phenazine pyocyanin as a highly potent prophage inducer of S. aureus that, in contrast to mitomycin C, displayed prophage selectivity. Mutagenesis and biochemical investigations confirm the existence of a noncanonical mechanism beyond SOS-response that is controlled by the intracellular oxidation level and is prophage-selective. Our results demonstrate that human pathogens can produce metabolites triggering lysogenic to lytic conversion in a prophage-selective manner. We anticipate our discovery to be the starting point of unveiling metabolite-mediated microbe-prophage interactions and laying the foundations for a selective small molecule controlled manipulation of prophage activity. These could be for example applied to control microbial communities by their built-in destruction mechanism in a novel form of phage therapy or for the construction of small molecule-inducible switches in synthetic biology.
Conflict of interest statement
The authors declare no competing financial interest.
Figures


Similar articles
-
Staphylococcus aureus Prophage-Encoded Protein Causes Abortive Infection and Provides Population Immunity against Kayviruses.mBio. 2023 Apr 25;14(2):e0249022. doi: 10.1128/mbio.02490-22. Epub 2023 Feb 13. mBio. 2023. PMID: 36779718 Free PMC article.
-
Diversity of Pseudomonas aeruginosa Temperate Phages.mSphere. 2022 Feb 23;7(1):e0101521. doi: 10.1128/msphere.01015-21. Epub 2022 Feb 23. mSphere. 2022. PMID: 35196122 Free PMC article.
-
ϕSa3mw Prophage as a Molecular Regulatory Switch of Staphylococcus aureus β-Toxin Production.J Bacteriol. 2019 Jun 21;201(14):e00766-18. doi: 10.1128/JB.00766-18. Print 2019 Jul 15. J Bacteriol. 2019. PMID: 30962356 Free PMC article.
-
Research progress of prophages.Yi Chuan. 2021 Mar 16;43(3):240-248. doi: 10.16288/j.yczz.20-355. Yi Chuan. 2021. PMID: 33724208 Review.
-
Yet another way that phage λ manipulates its Escherichia coli host: λrexB is involved in the lysogenic-lytic switch.Mol Microbiol. 2015 May;96(4):689-93. doi: 10.1111/mmi.12969. Epub 2015 Mar 16. Mol Microbiol. 2015. PMID: 25684601 Review.
Cited by
-
Viral diversity and host associations in microbial electrolysis cells.ISME Commun. 2024 Nov 15;4(1):ycae143. doi: 10.1093/ismeco/ycae143. eCollection 2024 Jan. ISME Commun. 2024. PMID: 39660013 Free PMC article.
-
Streptomyces secretes a siderophore that sensitizes competitor bacteria to phage infection.Nat Microbiol. 2025 Feb;10(2):362-373. doi: 10.1038/s41564-024-01910-8. Epub 2025 Jan 8. Nat Microbiol. 2025. PMID: 39779880
-
Bacterium secretes chemical inhibitor that sensitizes competitor to bacteriophage infection.bioRxiv [Preprint]. 2024 Jan 31:2024.01.31.578241. doi: 10.1101/2024.01.31.578241. bioRxiv. 2024. Update in: Nat Microbiol. 2025 Feb;10(2):362-373. doi: 10.1038/s41564-024-01910-8. PMID: 38352521 Free PMC article. Updated. Preprint.
-
Natural products influence bacteriophage infectivity.Nat Prod Rep. 2025 Aug 18. doi: 10.1039/d5np00014a. Online ahead of print. Nat Prod Rep. 2025. PMID: 40824115 Free PMC article. Review.
-
Historical contingencies and phage induction diversify bacterioplankton communities at the microscale.Proc Natl Acad Sci U S A. 2022 Jul 26;119(30):e2117748119. doi: 10.1073/pnas.2117748119. Epub 2022 Jul 21. Proc Natl Acad Sci U S A. 2022. PMID: 35862452 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

