Inhibition of Antiviral Innate Immunity by Foot-and-Mouth Disease Virus Lpro through Interaction with the N-Terminal Domain of Swine RNase L
- PMID: 33980594
- PMCID: PMC8274615
- DOI: 10.1128/JVI.00361-21
Inhibition of Antiviral Innate Immunity by Foot-and-Mouth Disease Virus Lpro through Interaction with the N-Terminal Domain of Swine RNase L
Abstract
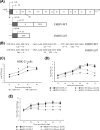
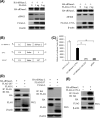
Foot-and-mouth disease virus (FMDV) is the pathogen of foot-and-mouth disease (FMD), which is a highly contagious disease in cloven-hoofed animals. To survive in the host, FMDV has evolved multiple strategies to antagonize host innate immune responses. In this study, we showed that the leader protease (Lpro) of FMDV, a papain-like proteinase, promoted viral replication by evading the antiviral interferon response through counteracting the 2',5'-oligoadenylate synthetase (OAS)/RNase L system. Specifically, we observed that the titers of Lpro deletion virus were significantly lower than those of wild-type FMDV (FMDV-WT) in cultured cells. Our mechanistic studies demonstrated that Lpro interfered with the OAS/RNase L pathway by interacting with the N-terminal domain of swine RNase L (sRNase L). Remarkably, Lpro of FMDV exhibited species-specific binding to RNase L in that the interaction was observed only in swine cells, not human, monkey, or canine cells. Lastly, we presented evidence that by interacting with sRNase L, FMDV Lpro inhibited cellular apoptosis. Taken together, these results demonstrate a novel mechanism that Lpro utilizes to escape the OAS/RNase L-mediated antiviral defense pathway. IMPORTANCE FMDV is a picornavirus that causes a significant disease in agricultural animals. FMDV has developed diverse strategies to escape the host interferon response. Here, we show that Lpro of FMDV antagonizes the OAS/RNase L pathway, an important interferon effector pathway, by interacting with the N-terminal domain of sRNase L. Interestingly, such a virus-host interaction is species-specific because the interaction is detected only in swine cells, not in human, monkey, or canine cells. Furthermore, Lpro inhibits apoptosis through interacting with sRNase L. This study demonstrates a novel mechanism by which FMDV has evolved to inhibit host innate immune responses.
Keywords: FMDV; ISGs; Lpro; RNase L; antagonistic mechanism.; foot-and-mouth disease virus.
Figures





Similar articles
-
Impairment of the DeISGylation Activity of Foot-and-Mouth Disease Virus Lpro Causes Attenuation In Vitro and In Vivo.J Virol. 2020 Jun 16;94(13):e00341-20. doi: 10.1128/JVI.00341-20. Print 2020 Jun 16. J Virol. 2020. PMID: 32295921 Free PMC article.
-
Foot-and-Mouth Disease Virus Antagonizes NOD2-Mediated Antiviral Effects by Inhibiting NOD2 Protein Expression.J Virol. 2019 May 15;93(11):e00124-19. doi: 10.1128/JVI.00124-19. Print 2019 Jun 1. J Virol. 2019. PMID: 30894473 Free PMC article.
-
Innate immune sensor LGP2 is cleaved by the Leader protease of foot-and-mouth disease virus.PLoS Pathog. 2018 Jun 29;14(6):e1007135. doi: 10.1371/journal.ppat.1007135. eCollection 2018 Jun. PLoS Pathog. 2018. PMID: 29958302 Free PMC article.
-
Multifunctional roles of leader protein of foot-and-mouth disease viruses in suppressing host antiviral responses.Vet Res. 2015 Oct 28;46:127. doi: 10.1186/s13567-015-0273-1. Vet Res. 2015. PMID: 26511922 Free PMC article. Review.
-
Evading the host immune response: how foot-and-mouth disease virus has become an effective pathogen.FEMS Immunol Med Microbiol. 2008 Jun;53(1):8-17. doi: 10.1111/j.1574-695X.2008.00409.x. Epub 2008 Apr 8. FEMS Immunol Med Microbiol. 2008. PMID: 18400012 Review.
Cited by
-
Reprogramming viral immune evasion for a rational design of next-generation vaccines for RNA viruses.Front Immunol. 2023 Apr 17;14:1172000. doi: 10.3389/fimmu.2023.1172000. eCollection 2023. Front Immunol. 2023. PMID: 37138878 Free PMC article. Review.
-
Picornavirus infection enhances aspartate by the SLC38A8 transporter to promote viral replication.PLoS Pathog. 2023 Feb 3;19(2):e1011126. doi: 10.1371/journal.ppat.1011126. eCollection 2023 Feb. PLoS Pathog. 2023. PMID: 36735752 Free PMC article.
-
Mutation of FMDV Lpro H138 residue drives viral attenuation in cell culture and in vivo in swine.Front Vet Sci. 2022 Oct 31;9:1028077. doi: 10.3389/fvets.2022.1028077. eCollection 2022. Front Vet Sci. 2022. PMID: 36387381 Free PMC article.
-
Rotavirus NSP1 Subverts the Antiviral Oligoadenylate Synthetase-RNase L Pathway by Inducing RNase L Degradation.mBio. 2022 Dec 20;13(6):e0299522. doi: 10.1128/mbio.02995-22. Epub 2022 Nov 22. mBio. 2022. PMID: 36413023 Free PMC article.
-
Foot-and-Mouth Disease Virus: Molecular Interplays with IFN Response and the Importance of the Model.Viruses. 2022 Sep 27;14(10):2129. doi: 10.3390/v14102129. Viruses. 2022. PMID: 36298684 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

