CD8+ T cell metabolism in infection and cancer
- PMID: 33981085
- PMCID: PMC8806153
- DOI: 10.1038/s41577-021-00537-8
CD8+ T cell metabolism in infection and cancer
Abstract
Cytotoxic CD8+ T cells play a key role in the elimination of intracellular infections and malignant cells and can provide long-term protective immunity. In the response to infection, CD8+ T cell metabolism is coupled to transcriptional, translational and epigenetic changes that are driven by extracellular metabolites and immunological signals. These programmes facilitate the adaptation of CD8+ T cells to the diverse and dynamic metabolic environments encountered in the circulation and in the tissues. In the setting of disease, both cell-intrinsic and cell-extrinsic metabolic cues contribute to CD8+ T cell dysfunction. In addition, changes in whole-body metabolism, whether through voluntary or disease-induced dietary alterations, can influence CD8+ T cell-mediated immunity. Defining the metabolic adaptations of CD8+ T cells in specific tissue environments informs our understanding of how these cells protect against pathogens and tumours and maintain tissue health at barrier sites. Here, we highlight recent findings revealing how metabolic networks enforce specific CD8+ T cell programmes and discuss how metabolism is integrated with CD8+ T cell differentiation and function and determined by environmental cues.
© 2021. Springer Nature Limited.
Conflict of interest statement
Competing Interests
M.R.-C. and N.E.S. declare no competing interests. A.W.G. serves on the scientific advisory boards of Pandion Therapeutics and Arsenal Bio.
Figures



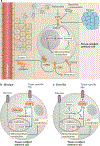

References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

