Identification of key genes and biological processes contributing to colitis associated dysplasia in ulcerative colitis
- PMID: 33987007
- PMCID: PMC8086577
- DOI: 10.7717/peerj.11321
Identification of key genes and biological processes contributing to colitis associated dysplasia in ulcerative colitis
Abstract
Background: Ulcerative colitis-associated colorectal cancer (UC-CRC) is a life-threatening complication of ulcerative colitis (UC). The mechanisms underlying UC-CRC remain to be elucidated. The purpose of this study was to explore the key genes and biological processes contributing to colitis-associated dysplasia (CAD) or carcinogenesis in UC via database mining, thus offering opportunities for early prediction and intervention of UC-CRC.
Methods: Microarray datasets (GSE47908 and GSE87466) were downloaded from Gene Expression Omnibus (GEO). Differentially expressed genes (DEGs) between groups of GSE47908 were identified using the "limma" R package. Weighted gene co-expression network analysis (WGCNA) based on DEGs between the CAD and control groups was conducted subsequently. Functional enrichment analysis was performed, and hub genes of selected modules were identified using the "clusterProfiler" R package. Single-gene gene set enrichment analysis (GSEA) was conducted to predict significant biological processes and pathways associated with the specified gene.
Results: Six functional modules were identified based on 4929 DEGs. Green and blue modules were selected because of their consistent correlation with UC and CAD, and the highest correlation coefficient with the progress of UC-associated carcinogenesis. Functional enrichment analysis revealed that genes of these two modules were significantly enriched in biological processes, including mitochondrial dysfunction, cell-cell junction, and immune responses. However, GSEA based on differential expression analysis between sporadic colorectal cancer (CRC) and normal controls from The Cancer Genome Atlas (TCGA) indicated that mitochondrial dysfunction may not be the major carcinogenic mechanism underlying sporadic CRC. Thirteen hub genes (SLC25A3, ACO2, AIFM1, ATP5A1, DLD, TFE3, UQCRC1, ADIPOR2, SLC35D1, TOR1AIP1, PRR5L, ATOX1, and DTX3) were identified. Their expression trends were validated in UC patients of GSE87466, and their potential carcinogenic effects in UC were supported by their known functions and other relevant studies reported in the literature. Single-gene GSEA indicated that biological processes and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways related to angiogenesis and immune response were positively correlated with the upregulation of TFE3, whereas those related to mitochondrial function and energy metabolism were negatively correlated with the upregulation of TFE3.
Conclusions: Using WGCNA, this study found two gene modules that were significantly correlated with CAD, of which 13 hub genes were identified as the potential key genes. The critical biological processes in which the genes of these two modules were significantly enriched include mitochondrial dysfunction, cell-cell junction, and immune responses. TFE3, a transcription factor related to mitochondrial function and cancers, may play a central role in UC-associated carcinogenesis.
Keywords: Colitis associated dysplasia; Ulcerative colitis; Ulcerative colitis associated colorectal cancer; Weighted gene co-expression network analysis.
©2021 Zhang et al.
Conflict of interest statement
The authors declare there are no competing interests.
Figures

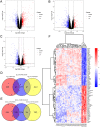





References
-
- Bruggemann M, Gromes A, Poss M, Schmidt D, Klumper N, Tolkach Y, Dietrich D, Kristiansen G, Muller SC, Ellinger J. Systematic analysis of the expression of the mitochondrial ATP synthase (Complex V) subunits in clear cell renal cell carcinoma. Translational Oncology. 2017;10:661–668. doi: 10.1016/j.tranon.2017.06.002. - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

