Sinorhizobium medicae WSM419 Genes That Improve Symbiosis between Sinorhizobium meliloti Rm1021 and Medicago truncatula Jemalong A17 and in Other Symbiosis Systems
- PMID: 33990306
- PMCID: PMC8276806
- DOI: 10.1128/AEM.03004-20
Sinorhizobium medicae WSM419 Genes That Improve Symbiosis between Sinorhizobium meliloti Rm1021 and Medicago truncatula Jemalong A17 and in Other Symbiosis Systems
Abstract
Some soil bacteria, called rhizobia, can interact symbiotically with legumes, in which they form nodules on the plant roots, where they can reduce atmospheric dinitrogen to ammonia, a form of nitrogen that can be used by growing plants. Rhizobium-plant combinations can differ in how successful this symbiosis is: for example, Sinorhizobium meliloti Rm1021 forms a relatively ineffective symbiosis with Medicago truncatula Jemalong A17, but Sinorhizobium medicae WSM419 is able to support more vigorous plant growth. Using proteomic data from free-living and symbiotic S. medicae WSM419, we previously identified a subset of proteins that were not closely related to any S. meliloti Rm1021 proteins and speculated that adding one or more of these proteins to S. meliloti Rm1021 would increase its effectiveness on M. truncatula A17. Three genes, Smed_3503, Smed_5985, and Smed_6456, were cloned into S. meliloti Rm1021 downstream of the E. coli lacZ promoter. Strains with these genes increased nodulation and improved plant growth, individually and in combination with one another. Smed_3503, renamed iseA (
Keywords: Medicago truncatula; Sinorhizobium medicae WSM419; Sinorhizobium meliloti Rm1021; alfalfa; autoregulation of nodulation (AON); model legume; nitrogen fixation; root nodules; symbiosis.
Figures
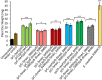
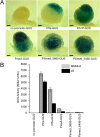
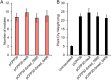
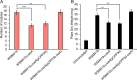
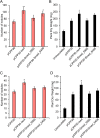
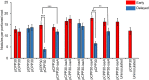
Similar articles
-
Nodule carbohydrate catabolism is enhanced in the Medicago truncatula A17-Sinorhizobium medicae WSM419 symbiosis.Front Microbiol. 2014 Aug 27;5:447. doi: 10.3389/fmicb.2014.00447. eCollection 2014. Front Microbiol. 2014. PMID: 25221545 Free PMC article.
-
Evidence that the exoH gene of Sinorhizobium meliloti does not appear to influence symbiotic effectiveness with Medicago truncatula 'Jemalong A17'.Can J Microbiol. 2010 Dec;56(12):996-1002. doi: 10.1139/W10-086. Can J Microbiol. 2010. PMID: 21164569
-
The model legume Medicago truncatula A17 is poorly matched for N2 fixation with the sequenced microsymbiont Sinorhizobium meliloti 1021.New Phytol. 2008;179(1):62-66. doi: 10.1111/j.1469-8137.2008.02464.x. Epub 2008 Apr 14. New Phytol. 2008. PMID: 18422896
-
Exploring the role of symbiotic modifier peptidases in the legume - rhizobium symbiosis.Arch Microbiol. 2024 Mar 11;206(4):147. doi: 10.1007/s00203-024-03920-w. Arch Microbiol. 2024. PMID: 38462552 Review.
-
How rhizobial symbionts invade plants: the Sinorhizobium-Medicago model.Nat Rev Microbiol. 2007 Aug;5(8):619-33. doi: 10.1038/nrmicro1705. Nat Rev Microbiol. 2007. PMID: 17632573 Free PMC article. Review.
Cited by
-
Horizontal gene transfer of the Mer operon is associated with large effects on the transcriptome and increased tolerance to mercury in nitrogen-fixing bacteria.BMC Microbiol. 2024 Jul 6;24(1):247. doi: 10.1186/s12866-024-03391-5. BMC Microbiol. 2024. PMID: 38971740 Free PMC article.
-
A protease and a lipoprotein jointly modulate the conserved ExoR-ExoS-ChvI signaling pathway critical in Sinorhizobium meliloti for symbiosis with legume hosts.PLoS Genet. 2023 Oct 23;19(10):e1010776. doi: 10.1371/journal.pgen.1010776. eCollection 2023 Oct. PLoS Genet. 2023. PMID: 37871041 Free PMC article.
-
A minimized symbiotic gene set from the 1.68 Mb pSymB chromid of Sinorhizobium meliloti reveals auxiliary symbiotic loci.BMC Biol. 2025 Jul 9;23(1):204. doi: 10.1186/s12915-025-02298-5. BMC Biol. 2025. PMID: 40629387 Free PMC article.
References
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Other Literature Sources

