100-My history of bornavirus infections hidden in vertebrate genomes
- PMID: 33990470
- PMCID: PMC8157955
- DOI: 10.1073/pnas.2026235118
100-My history of bornavirus infections hidden in vertebrate genomes
Abstract
Although viruses have threatened our ancestors for millions of years, prehistoric epidemics of viruses are largely unknown. Endogenous bornavirus-like elements (EBLs) are ancient bornavirus sequences derived from the viral messenger RNAs that were reverse transcribed and inserted into animal genomes, most likely by retrotransposons. These elements can be used as molecular fossil records to trace past bornaviral infections. In this study, we systematically identified EBLs in vertebrate genomes and revealed the history of bornavirus infections over nearly 100 My. We confirmed that ancient bornaviral infections have occurred in diverse vertebrate lineages, especially in primate ancestors. Phylogenetic analyses indicated that primate ancestors were infected with various bornaviral lineages during evolution. EBLs in primate genomes formed clades according to their integration ages, suggesting that bornavirus lineages infected with primate ancestors had changed chronologically. However, some bornaviral lineages may have coexisted with primate ancestors and underwent repeated endogenizations for tens of millions of years. Moreover, a bornaviral lineage that coexisted with primate ancestors also endogenized in the genomes of some ancestral bats. The habitats of these bat ancestors have been reported to overlap with the migration route of primate ancestors. These results suggest that long-term virus-host coexistence expanded the geographic distributions of the bornaviral lineage along with primate migration and may have spread their infections to these bat ancestors. Our findings provide insight into the history of bornavirus infections over geological timescales that cannot be deduced from research using extant viruses alone, thus broadening our perspective on virus-host coevolution.
Keywords: ancient viral infection; endogenous bornavirus-like element; paleovirology; vertebrate evolution; virus–host coevolutionary history.
Conflict of interest statement
The authors declare no competing interest.
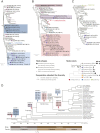
Figures


Similar articles
-
Systematic estimation of insertion dates of endogenous bornavirus-like elements in vesper bats.J Vet Med Sci. 2018 Aug 30;80(8):1356-1363. doi: 10.1292/jvms.18-0211. Epub 2018 Jul 5. J Vet Med Sci. 2018. PMID: 29973433 Free PMC article.
-
Divergent bornaviruses from Australian carpet pythons with neurological disease date the origin of extant Bornaviridae prior to the end-Cretaceous extinction.PLoS Pathog. 2018 Feb 20;14(2):e1006881. doi: 10.1371/journal.ppat.1006881. eCollection 2018 Feb. PLoS Pathog. 2018. PMID: 29462190 Free PMC article.
-
Genomic Mining Reveals Deep Evolutionary Relationships between Bornaviruses and Bats.Viruses. 2015 Nov 10;7(11):5792-800. doi: 10.3390/v7112906. Viruses. 2015. PMID: 26569285 Free PMC article.
-
Paleovirology of bornaviruses: What can be learned from molecular fossils of bornaviruses.Virus Res. 2019 Mar;262:2-9. doi: 10.1016/j.virusres.2018.04.006. Epub 2018 Apr 6. Virus Res. 2019. PMID: 29630909 Review.
-
Comprehensive analysis of endogenous bornavirus-like elements in eukaryote genomes.Philos Trans R Soc Lond B Biol Sci. 2013 Aug 12;368(1626):20120499. doi: 10.1098/rstb.2012.0499. Print 2013 Sep 19. Philos Trans R Soc Lond B Biol Sci. 2013. PMID: 23938751 Free PMC article. Review.
Cited by
-
Historic and Prehistoric Epidemics: An Overview of Sources Available for the Study of Ancient Pathogens.Epidemiologia (Basel). 2022 Oct 7;3(4):443-464. doi: 10.3390/epidemiologia3040034. Epidemiologia (Basel). 2022. PMID: 36547255 Free PMC article. Review.
-
Genomic transfers help to decipher the ancient evolution of filoviruses and interactions with vertebrate hosts.PLoS Pathog. 2024 Sep 3;20(9):e1011864. doi: 10.1371/journal.ppat.1011864. eCollection 2024 Sep. PLoS Pathog. 2024. PMID: 39226335 Free PMC article.
-
Current trends in RNA virus detection through metatranscriptome sequencing data.FEBS Open Bio. 2023 Jun;13(6):992-1000. doi: 10.1002/2211-5463.13626. Epub 2023 May 20. FEBS Open Bio. 2023. PMID: 37163224 Free PMC article. Review.
-
Endogenous nege-like viral elements in arthropod genomes reveal virus-host coevolution and ancient history of two plant virus families.J Virol. 2024 Oct 22;98(10):e0099724. doi: 10.1128/jvi.00997-24. Epub 2024 Aug 30. J Virol. 2024. PMID: 39212930 Free PMC article.
-
The hidden diversity of ancient bornaviral sequences from X and P genes in vertebrate genomes.Virus Evol. 2023 Jun 3;9(1):vead038. doi: 10.1093/ve/vead038. eCollection 2023. Virus Evol. 2023. PMID: 37360682 Free PMC article.
References
-
- Taubenberger J. K., Reid A. H., Krafft A. E., Bijwaard K. E., Fanning T. G., Initial genetic characterization of the 1918 “Spanish” influenza virus. Science 275, 1793–1796 (1997). - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

