lncRNA MALAT1 Promotes Renal Fibrosis in Diabetic Nephropathy by Targeting the miR-2355-3p/IL6ST Axis
- PMID: 33995063
- PMCID: PMC8117091
- DOI: 10.3389/fphar.2021.647650
lncRNA MALAT1 Promotes Renal Fibrosis in Diabetic Nephropathy by Targeting the miR-2355-3p/IL6ST Axis
Abstract
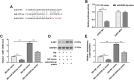
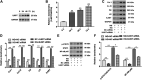
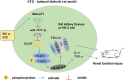
Long noncoding RNA (lncRNAs) metastasis-associated lung adenocarcinoma transcript 1 (MALAT1) has been reported in diabetic nephropathy (DN) about its effect on podocyte function and cell heat shock induced by hyperglycemia. However, the biological mechanism of MALAT1 regulating DN fibrosis needs further study. In this study, SD rats were administrated with streptozotocin (STZ) to establish a diabetes model. In vitro, human renal tubular epithelial cells (HK-2 and 293T) were treated with high glucose (HG). Here, we found that MALAT1 was upregulated in renal tissues of diabetic rats and HG-treated cells, and HG treatment promoted cell proliferation and invasion. MALAT1 overexpression aggravated protein levels of collagen I (col I), collagen IV (col IV), fibronectin (FN), and laminin (LN) in HK-2 cells, while MALAT1 knockdown exerted the opposite effect. Moreover, the luciferase reporter gene and pull-down assays demonstrated that MALAT1 interacted with miR-2355-3p. The miR-2355-3p level was downregulated in diabetic rats and HG-treated cells, and MALAT1 overexpression inhibited the miR-2355-3p level. Bioinformatics prediction and luciferase reporter gene assay revealed that interleukin 6 signal transducer (IL6ST) was a target of miR-2355-3p. In addition, miR-2355-3p overexpression attenuated fibrosis-related gene levels in HG-treated cells by inhibiting IL6ST expression and inactivating the recombinant signal transducer and activator of the transcription 3 (STAT3) signaling pathway. Knockdown of miR-2355-3p reversed the inhibitory effect of MALAT1 knockdown on IL6ST, col I, col IV, FN, and LN protein levels in HG-induced cells. Overexpression of MALAT1 aggravated cell damage in HG-induced cells via the miR-2355-3p/IL6ST/STAT3 signaling pathway. Finally, enhanced renal fibrosis and kidney tissue damage were observed in diabetic rats. In conclusion, MALAT1 overexpression may enhance renal fibrosis in diabetic rats and cell damage in HG-induced HK-2 cells via the miR-2355-3p/IL6ST axis, which provides a new perspective of DN treatment.
Keywords: IL6ST; STAT3 pathway; diabetic nephropathy; lncRNA MALAT1; miR-2355-3p.
Copyright © 2021 Huang, Zhang and Ge.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures






References
-
- Amanzadeh M., Mota A., Zarghami N., Abedi-Azar S., Abroon S., Akbarian N., et al. (2018). Association between matrix metalloproteinase-3 activity and glomerular filtration rate and albuminuria status in patients with type 2 diabetes mellitus. Iran. J. Kidney Dis. 12, 40–47. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

