Recording and Simulating Proton-Related Metabolism in Bacterial Cell Suspensions
- PMID: 33995312
- PMCID: PMC8117226
- DOI: 10.3389/fmicb.2021.654065
Recording and Simulating Proton-Related Metabolism in Bacterial Cell Suspensions
Abstract
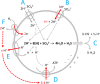
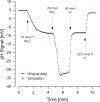
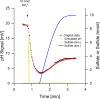
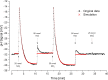
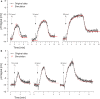
Proton release and uptake induced by metabolic activities were measured in non-buffered cell suspensions by means of a pH electrode. Recorded data were used for simulating substrate turnover rates by means of a new freeware app (proton.exe). The program applies Michaelis-Menten or first-order kinetics to the metabolic processes and allows for parametrization of simultaneously ongoing processes. The simulation includes changes of the transmembrane ΔpH, membrane potential and ATP gains, and demonstrates the principles of chemiosmotic energy conservation. In our experiments, the versatile sulfate-reducing bacterium Desulfovibrio desulfuricans CSN (DSM 9104) was used as model organism. We analysed sulfate uptake by proton-sulfate symport, scalar alkalinization by sulfate reduction to sulfide, as well as nitrate and nitrite reduction to ammonia, and electron transport-coupled proton translocation. Two types of experiments were performed: In oxidant pulse experiments, cells were kept under H2, and micromolar amounts of sulfate, nitrate or nitrite were added. For reductant pulse experiments, small amounts of H2-saturated KCl were added to cells incubated under N2 with an excess of one of the above-mentioned electron acceptors. To study electron-transport driven proton translocation, the membrane potential was neutralized by addition of KSCN (100 mM). H+/e- ratios of electron-transport driven proton translocation were calculated by simulation with proton.exe. This method gave lower but more realistic values than logarithmic extrapolation. We could verify the kinetic simulation parameters found with proton.exe using series of increasing additions of the reactants. Our approach allows for studying a broad variety of proton-related metabolic activities at micromolar concentrations and time scales of seconds to minutes.
Keywords: ATP synthase activity; Desulfovibrio desulfuricans; Michaelis-Menten kinetics; dissimilatory nitrate reduction to ammonia; dissimilatory sulfate reduction; proton-sulfate symport; vectorial proton translocation.
Copyright © 2021 Cypionka and Reese.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
Similar articles
-
Bioenergetics of the alkaliphilic sulfate-reducing bacterium Desulfonatronovibrio hydrogenovorans.Microbiology (Reading). 2002 Mar;148(Pt 3):853-860. doi: 10.1099/00221287-148-3-853. Microbiology (Reading). 2002. PMID: 11882721
-
Understanding the response of Desulfovibrio desulfuricans ATCC 27774 to the electron acceptors nitrate and sulfate - biosynthetic costs modulate substrate selection.Biochim Biophys Acta Proteins Proteom. 2017 Nov;1865(11 Pt A):1455-1469. doi: 10.1016/j.bbapap.2017.07.021. Epub 2017 Aug 25. Biochim Biophys Acta Proteins Proteom. 2017. PMID: 28847524
-
Characterization of sulfate transport in Desulfovibrio desulfuricans.Arch Microbiol. 1989;152(3):237-43. doi: 10.1007/BF00409657. Arch Microbiol. 1989. PMID: 2476099
-
Oxygen respiration by desulfovibrio species.Annu Rev Microbiol. 2000;54:827-48. doi: 10.1146/annurev.micro.54.1.827. Annu Rev Microbiol. 2000. PMID: 11018146 Review.
-
Evolution of membrane bioenergetics.J Supramol Struct. 1980;13(4):421-46. doi: 10.1002/jss.400130403. J Supramol Struct. 1980. PMID: 6453255 Review.
Cited by
-
Multi-Modal Multi-Array Electrochemical and Optical Sensor Suite for a Biological CubeSat Payload.Sensors (Basel). 2024 Jan 2;24(1):265. doi: 10.3390/s24010265. Sensors (Basel). 2024. PMID: 38203127 Free PMC article.
References
-
- Badziong W., Thauer R. K. (1980) Vectorial electron transport in Desulfovibrio vulgaris (Marburg) growing on hydrogen plus sulfate as sole energy source. Arch. Microbiol. 125, 167–174. 10.1007/BF00403215 - DOI
-
- Crozier T. E., Yamamoto S. (1974). Solubility of hydrogen in water, sea water, and sodium chloridesolutions. J. Chem. Eng. Data 19 242–244. 10.1021/je60062a007 - DOI
-
- Cypionka H. (1986). Sulfide-controlled continuous culture of sulfate-reducing bacteria. J. Microbiol. Meth. 5 1–9. 10.1016/0167-7012(86)90018-7 - DOI
-
- Cypionka H. (1987). Uptake of sulfate, sulfite and thiosulfate by proton-anion symport in Desulfovibrio desulfuricans. Arch. Microbiol. 148 144–149. 10.1007/bf00425363 - DOI
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

